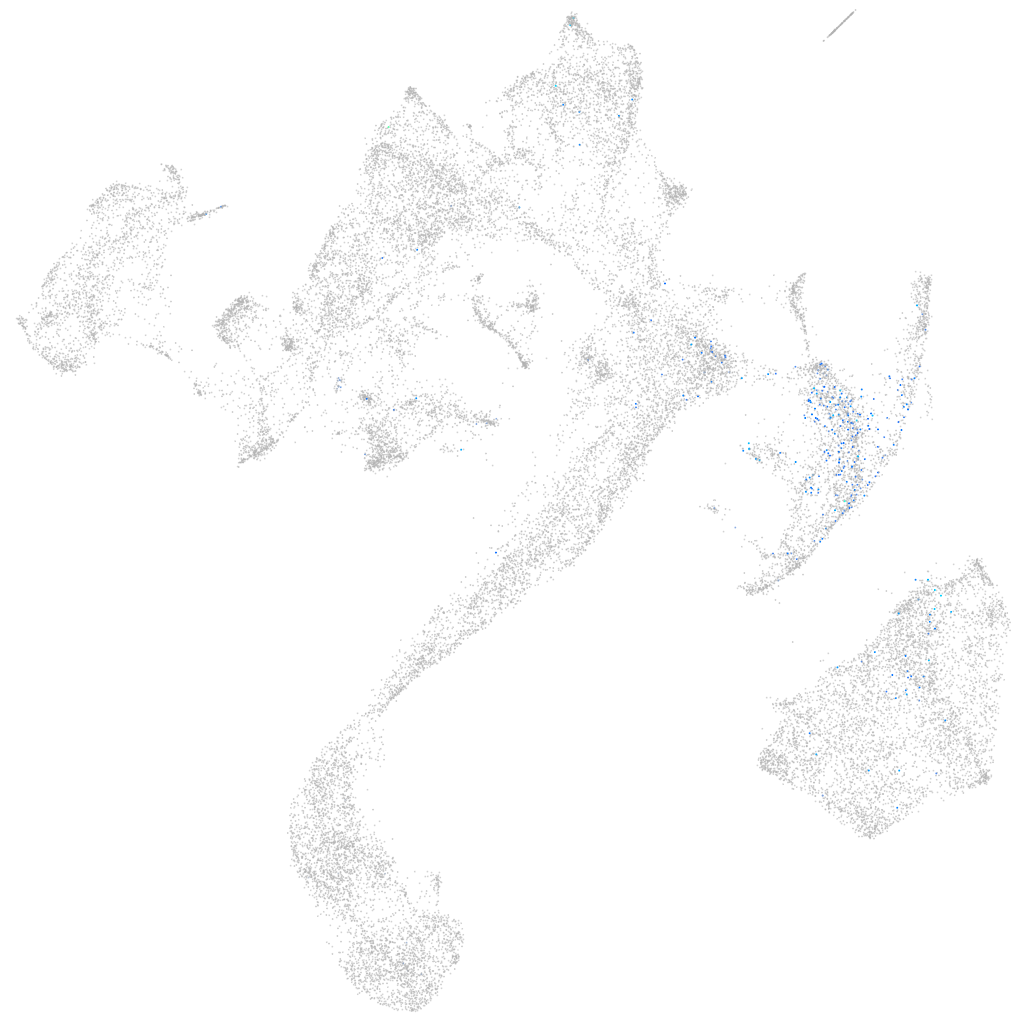
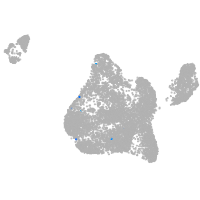
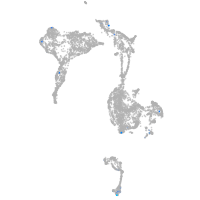
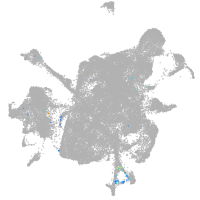
solute carrier family 6 member 7
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| pcdh8 | 0.165 | fabp3 | -0.075 |
| her1 | 0.163 | actc1b | -0.063 |
| grin2da | 0.142 | pabpc4 | -0.058 |
| tbx6 | 0.141 | ak1 | -0.056 |
| itm2cb | 0.135 | atp2a1 | -0.056 |
| her7 | 0.134 | sparc | -0.055 |
| msgn1 | 0.130 | ckma | -0.055 |
| myf5 | 0.126 | ckmb | -0.055 |
| fn1b | 0.125 | tmem38a | -0.055 |
| apoc1 | 0.121 | bhmt | -0.054 |
| nid2a | 0.120 | gamt | -0.054 |
| hoxb10a | 0.120 | eno1a | -0.054 |
| hoxd10a | 0.118 | ttn.2 | -0.054 |
| draxin | 0.118 | mylpfa | -0.052 |
| her11 | 0.115 | aldoab | -0.052 |
| BX005254.3 | 0.112 | gapdh | -0.052 |
| hes6 | 0.112 | eef1da | -0.050 |
| dlc | 0.112 | srl | -0.050 |
| hoxc3a | 0.111 | acta1b | -0.050 |
| tbx16l | 0.111 | neb | -0.050 |
| LO016987.2 | 0.110 | rplp2 | -0.050 |
| ripply2 | 0.109 | plecb | -0.049 |
| gnaia | 0.108 | myl1 | -0.049 |
| hoxb7a | 0.107 | tnnc2 | -0.049 |
| her12 | 0.106 | cox17 | -0.049 |
| ephb3 | 0.106 | cav3 | -0.048 |
| phc2a | 0.106 | ldb3b | -0.048 |
| ism1 | 0.103 | tnnt3a | -0.048 |
| rem1 | 0.102 | ndrg2 | -0.048 |
| tspan7 | 0.101 | atp1a2a | -0.047 |
| greb1 | 0.100 | CABZ01078594.1 | -0.047 |
| alpi.1 | 0.099 | nme2b.2 | -0.047 |
| kcnj19a | 0.096 | si:dkey-16p21.8 | -0.047 |
| cx43.4 | 0.094 | ldb3a | -0.047 |
| BX001014.2 | 0.094 | mylz3 | -0.047 |