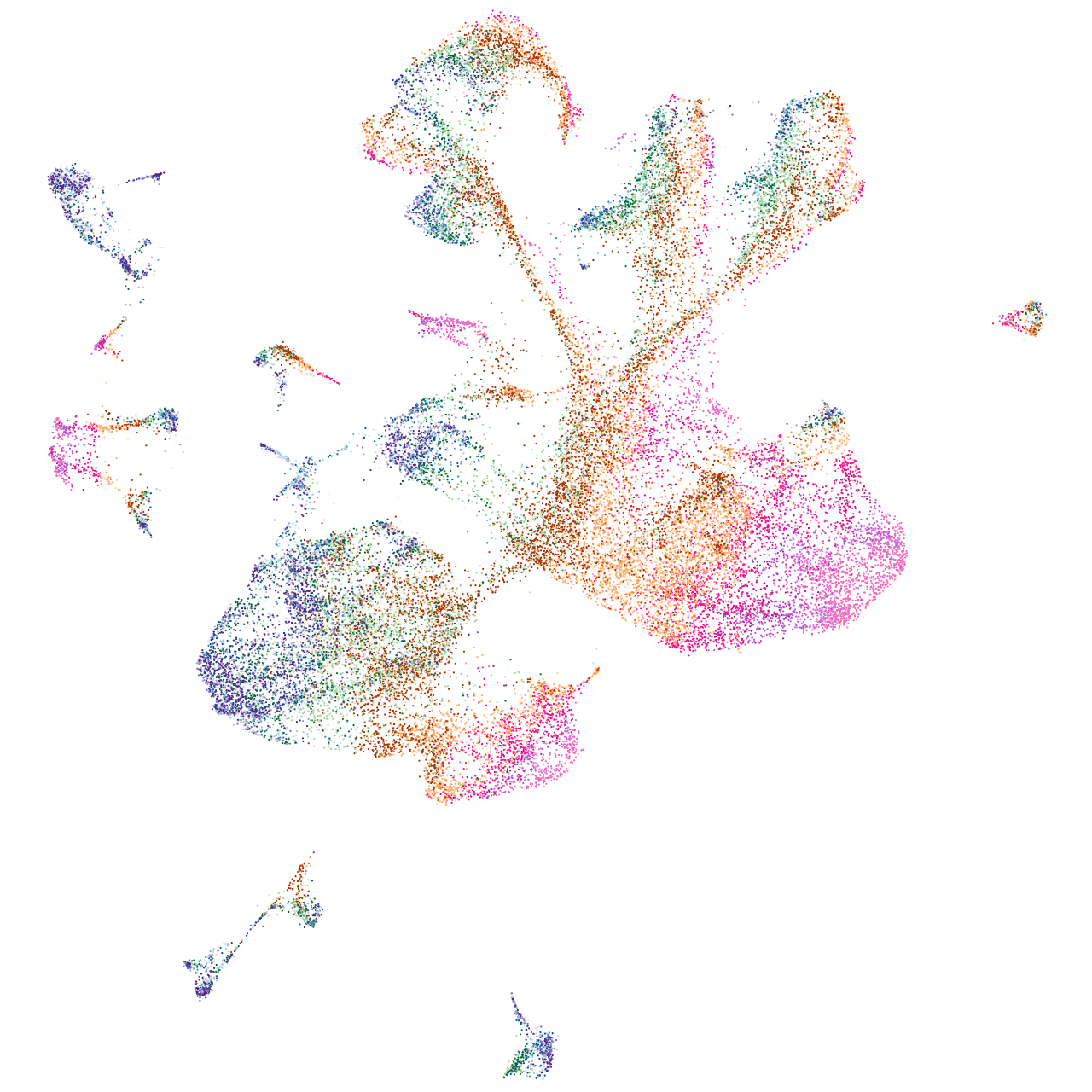
Sp5 transcription factor-like
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| noto | 0.499 | fabp3 | -0.180 |
| fgf8a | 0.479 | gpm6aa | -0.172 |
| cdx4 | 0.476 | tmsb4x | -0.165 |
| nradd | 0.468 | CR383676.1 | -0.163 |
| pou5f3 | 0.437 | marcksl1b | -0.162 |
| lrrc17 | 0.418 | ckbb | -0.150 |
| XLOC-042222 | 0.415 | nova2 | -0.147 |
| nkx1.2la | 0.406 | fabp7a | -0.139 |
| hspb1 | 0.387 | rtn1a | -0.130 |
| dnase1l4.1 | 0.378 | pvalb1 | -0.126 |
| apela | 0.372 | pvalb2 | -0.123 |
| apoc1 | 0.344 | meis1b | -0.121 |
| chrd | 0.337 | CU467822.1 | -0.119 |
| BX005254.3 | 0.334 | hbbe1.3 | -0.115 |
| cpm | 0.327 | COX3 | -0.114 |
| tbxta | 0.313 | tmsb | -0.114 |
| cxcr4a | 0.309 | actc1b | -0.114 |
| foxd5 | 0.308 | zgc:165461 | -0.113 |
| znfl2a | 0.304 | gpm6ab | -0.108 |
| nid2a | 0.300 | vim | -0.108 |
| cdx1a | 0.298 | hbae3 | -0.107 |
| BX927258.1 | 0.295 | atp1a1b | -0.106 |
| eve1 | 0.289 | gapdhs | -0.104 |
| BX001014.2 | 0.280 | nr2f1a | -0.104 |
| hoxb10a | 0.276 | CU634008.1 | -0.102 |
| pnx | 0.274 | stmn1b | -0.101 |
| npm1a | 0.272 | gpm6bb | -0.101 |
| sall4 | 0.271 | rnasekb | -0.100 |
| her3 | 0.270 | marcksl1a | -0.098 |
| hnf1ba | 0.269 | myt1b | -0.097 |
| mllt3 | 0.267 | slc1a2b | -0.097 |
| hoxd10a | 0.267 | cadm3 | -0.096 |
| znfl1k | 0.262 | gnao1a | -0.095 |
| arid3b | 0.261 | ppdpfb | -0.095 |
| hoxd12a | 0.260 | si:dkeyp-75h12.5 | -0.094 |