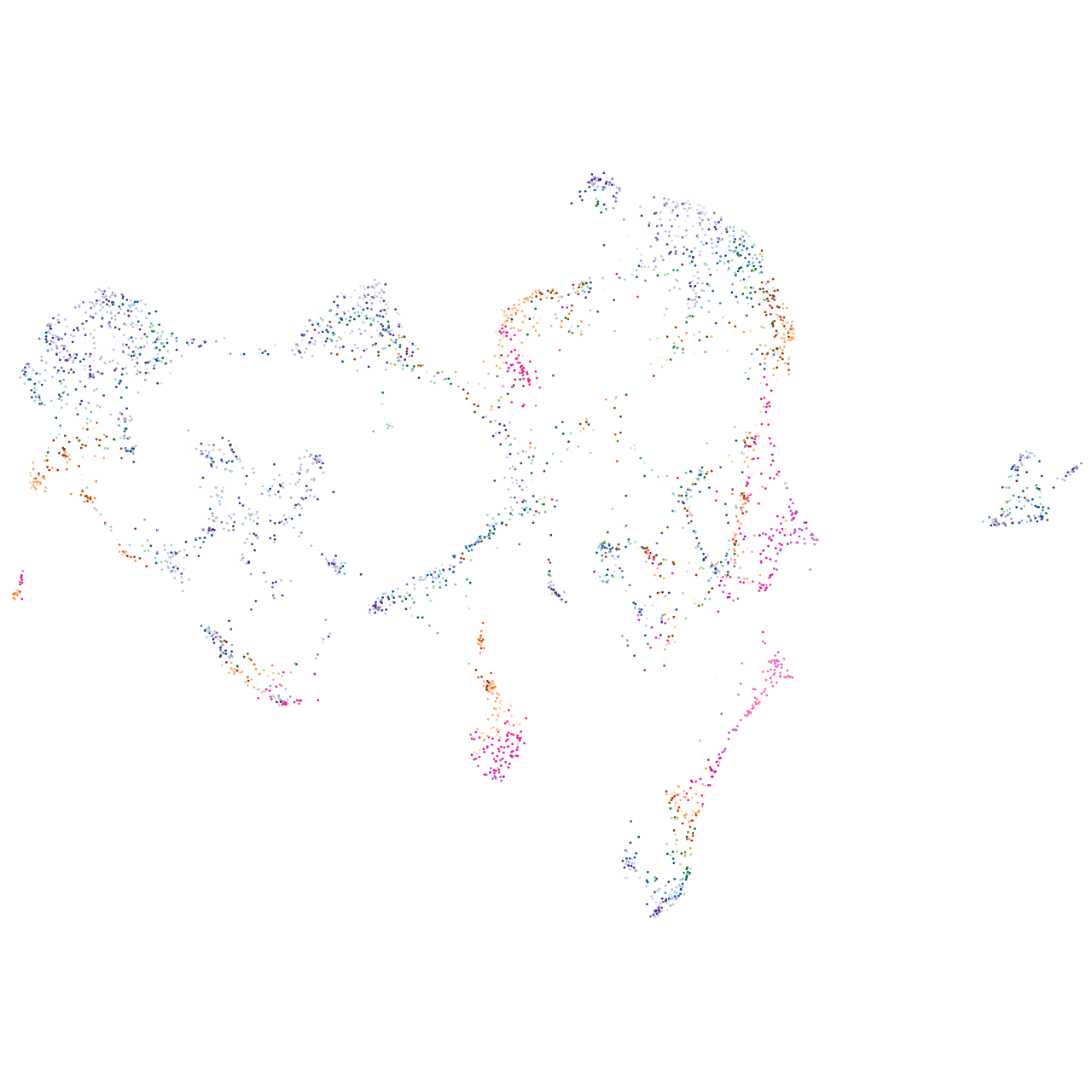
"protein kinase, AMP-activated, alpha 1 catalytic subunit"
ZFIN 
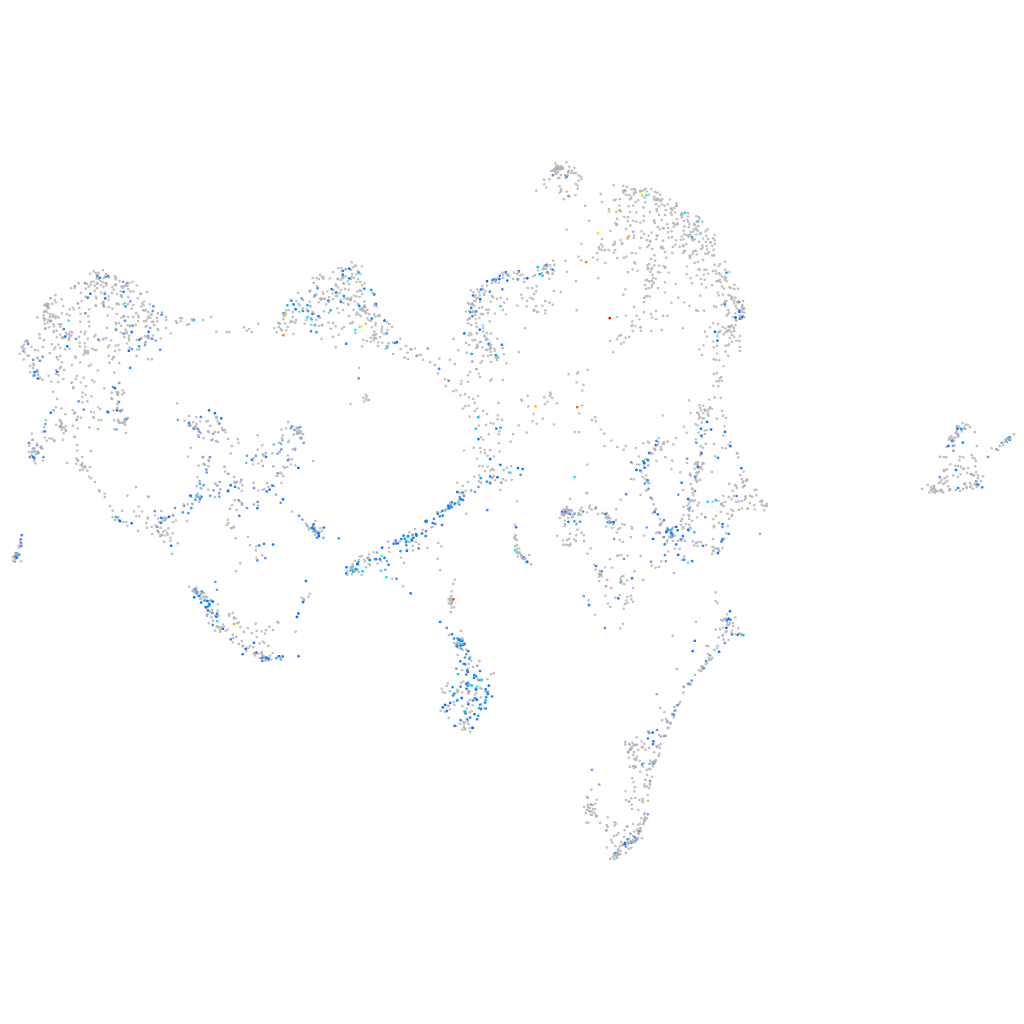
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| slc41a1 | 0.277 | agr2 | -0.167 |
| atp6v0ca | 0.263 | si:dkey-65b12.6 | -0.146 |
| atp6v1g1 | 0.255 | krt91 | -0.144 |
| atp6v1e1b | 0.255 | muc5.1 | -0.142 |
| foxi3a | 0.253 | krt5 | -0.137 |
| etf1a | 0.253 | spaca4l | -0.136 |
| atp6v1aa | 0.252 | b3gnt7 | -0.126 |
| atp6v1c1b | 0.250 | SPDEF | -0.124 |
| atp6v0a1a | 0.250 | nansb | -0.124 |
| slc6a6b | 0.247 | atp2a3 | -0.123 |
| atp6v1ba | 0.245 | galnt7 | -0.122 |
| gcm2 | 0.244 | zgc:112994 | -0.121 |
| ca15a | 0.243 | sytl2b | -0.119 |
| atp6ap1b | 0.242 | foxa3 | -0.117 |
| atp5mc1 | 0.240 | LOC110438378 | -0.117 |
| rnaseka | 0.236 | fxyd1 | -0.117 |
| ceacam1 | 0.233 | cebpd | -0.116 |
| nipsnap2 | 0.231 | kita | -0.115 |
| uqcrc1 | 0.231 | p2rx1 | -0.113 |
| rpia | 0.230 | rpz5 | -0.111 |
| atp6v1h | 0.229 | cd63 | -0.111 |
| ca2 | 0.229 | zgc:92380 | -0.110 |
| atp6ap2 | 0.229 | tcima | -0.109 |
| atp6v0b | 0.229 | stard10 | -0.108 |
| atp6v0d1 | 0.228 | muc5.2 | -0.107 |
| sucla2 | 0.228 | kdelr3 | -0.106 |
| ndufb7 | 0.228 | gng13b | -0.106 |
| rasgrp4 | 0.227 | mir375-2 | -0.105 |
| tfe3b | 0.227 | kdelr2b | -0.104 |
| idh2 | 0.226 | pou2f3 | -0.103 |
| slc4a1b | 0.224 | galnt12 | -0.102 |
| si:dkey-192d15.2 | 0.223 | crb3b | -0.102 |
| uqcrfs1 | 0.223 | CABZ01075242.1 | -0.101 |
| atp5l | 0.220 | ier2b | -0.100 |
| cox7a2a | 0.220 | klf3 | -0.100 |