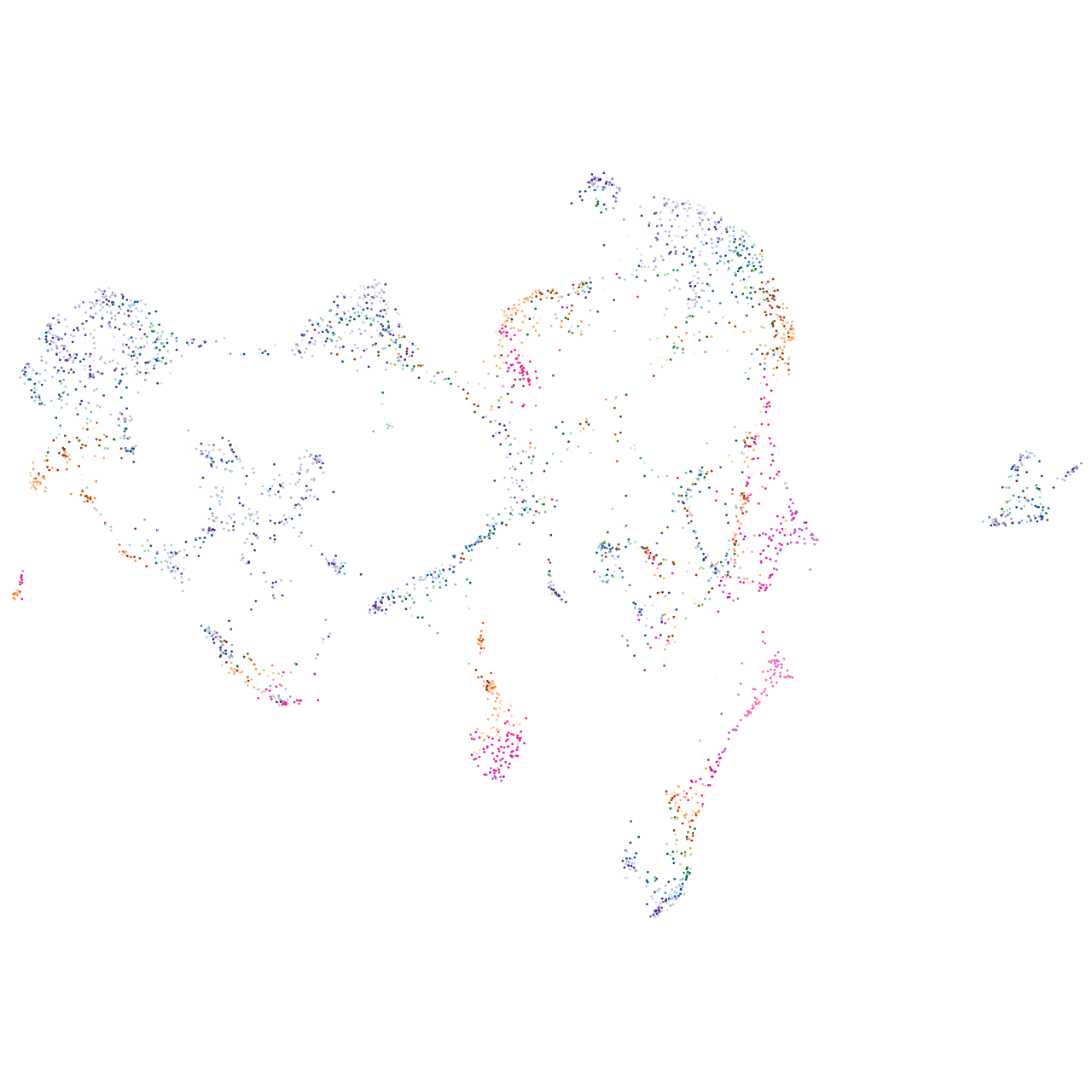
myelin basic protein b
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| cotl1 | 0.280 | agr2 | -0.152 |
| ca15a | 0.255 | muc5.1 | -0.133 |
| actn1 | 0.241 | foxa3 | -0.131 |
| atp6v0a1a | 0.241 | si:dkey-65b12.6 | -0.131 |
| myl12.1 | 0.239 | kita | -0.130 |
| atp6v1g1 | 0.234 | rpl38 | -0.129 |
| atp6ap2 | 0.231 | stard10 | -0.127 |
| arg2 | 0.230 | sytl2b | -0.123 |
| sult2st1 | 0.229 | SPDEF | -0.116 |
| csrp1a | 0.227 | cers3b | -0.116 |
| rac2 | 0.223 | BX000438.2 | -0.116 |
| slc31a2 | 0.222 | pou2f3 | -0.115 |
| ceacam1 | 0.222 | gng13b | -0.114 |
| ca2 | 0.222 | atp2a3 | -0.114 |
| zgc:110339 | 0.220 | tspan13a | -0.113 |
| zgc:56622 | 0.219 | b3gnt7 | -0.112 |
| actb2 | 0.219 | zgc:112994 | -0.110 |
| rpia | 0.219 | nansb | -0.107 |
| pfas | 0.219 | rps29 | -0.107 |
| atp6ap1b | 0.218 | si:dkey-151g10.6 | -0.107 |
| naga | 0.217 | si:dkey-87o1.2 | -0.105 |
| atp6v1e1b | 0.212 | mir375-2 | -0.105 |
| scgn | 0.210 | p2rx1 | -0.104 |
| cldni | 0.208 | fev | -0.103 |
| atp6v1aa | 0.206 | muc5.2 | -0.101 |
| degs1 | 0.203 | galnt7 | -0.100 |
| atp6v1h | 0.203 | rps21 | -0.100 |
| tp63 | 0.202 | rpl36 | -0.098 |
| atp6v1d | 0.201 | itpr3 | -0.096 |
| cebpb | 0.201 | rpl37 | -0.096 |
| rnaseka | 0.200 | LOC103910015 | -0.094 |
| atp6v1c1b | 0.200 | calm1b | -0.094 |
| sh3d21 | 0.200 | pdia5 | -0.094 |
| itgb1b.1 | 0.200 | foxa1 | -0.092 |
| cyp3c1 | 0.198 | slc46a3 | -0.092 |