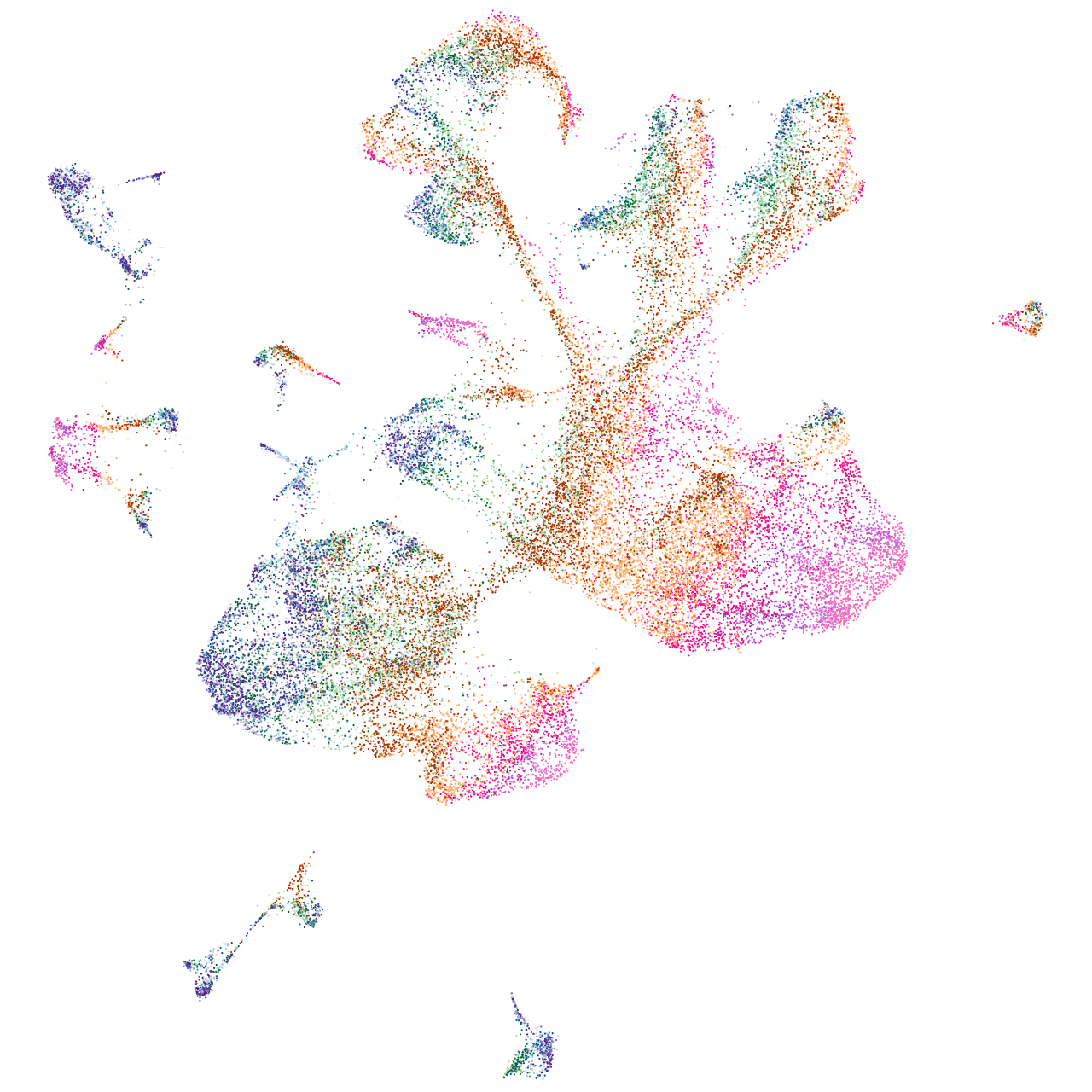
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| CU467822.1 | 0.085 | CABZ01075068.1 | -0.052 |
| ppap2d | 0.077 | lin28a | -0.043 |
| CR848047.1 | 0.075 | NC-002333.4 | -0.042 |
| fabp7a | 0.075 | onecut1 | -0.037 |
| efhd1 | 0.075 | onecut2 | -0.036 |
| gpm6bb | 0.074 | trim71 | -0.036 |
| cspg5b | 0.074 | si:ch211-222l21.1 | -0.036 |
| atp1b4 | 0.072 | snap25a | -0.035 |
| slc3a2a | 0.070 | elavl3 | -0.035 |
| slc6a11b | 0.069 | fthl27 | -0.034 |
| mfge8a | 0.067 | aplp1 | -0.031 |
| zgc:165461 | 0.067 | syt2a | -0.030 |
| cx43 | 0.066 | oc90 | -0.030 |
| slc1a2b | 0.066 | NC-002333.17 | -0.030 |
| fjx1 | 0.066 | isl1 | -0.030 |
| slc6a1b | 0.065 | nr6a1a | -0.030 |
| eno1b | 0.064 | sncb | -0.030 |
| gnai2a | 0.064 | ptmab | -0.030 |
| sept8b | 0.063 | isl2a | -0.030 |
| si:ch211-66e2.5 | 0.063 | stxbp1a | -0.029 |
| slc4a4a | 0.062 | h3f3d | -0.029 |
| GCA | 0.062 | elavl4 | -0.029 |
| cd63 | 0.061 | hoxc3a | -0.029 |
| acadl | 0.060 | stx1b | -0.029 |
| qki2 | 0.060 | pabpc1a | -0.028 |
| si:ch1073-303k11.2 | 0.059 | greb1l | -0.028 |
| prdx2 | 0.059 | zgc:100920 | -0.028 |
| nck1a | 0.059 | hoxb1b | -0.028 |
| CU634008.1 | 0.059 | slit3 | -0.028 |
| glula | 0.059 | dhrs3b | -0.028 |
| si:ch211-286b5.5 | 0.058 | gng2 | -0.028 |
| spry2 | 0.057 | kcnc3a | -0.027 |
| s1pr1 | 0.057 | gng3 | -0.027 |
| zgc:162780 | 0.057 | CABZ01072614.1 | -0.027 |
| gpr37l1b | 0.057 | cygb1 | -0.027 |