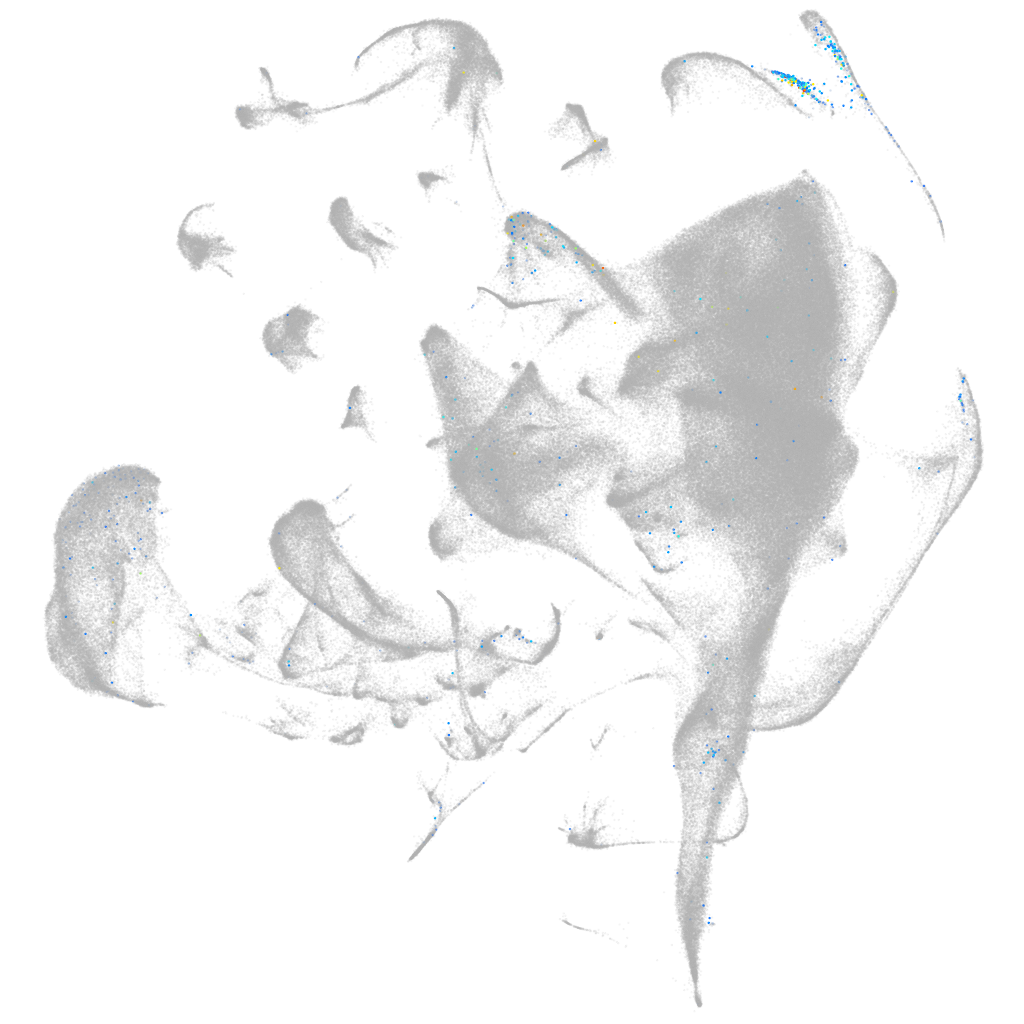
"obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF a"
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| tnni2a.3 | 0.221 | gpm6aa | -0.024 |
| tnni2a.1 | 0.218 | marcksl1a | -0.022 |
| fhl5 | 0.207 | hmgb1b | -0.021 |
| mybpc1 | 0.201 | mdkb | -0.021 |
| lmod2b | 0.192 | si:ch211-288g17.3 | -0.021 |
| myhb | 0.184 | elavl3 | -0.020 |
| klhl41a | 0.174 | gpm6ab | -0.020 |
| fhl1a | 0.161 | chd4a | -0.018 |
| ryr1a | 0.161 | cspg5a | -0.018 |
| myl4 | 0.160 | nova2 | -0.018 |
| sync | 0.160 | rtn1b | -0.018 |
| zgc:172270 | 0.158 | stmn1b | -0.018 |
| flnca | 0.156 | tmeff1b | -0.018 |
| AGBL1 | 0.154 | zc4h2 | -0.018 |
| mustn1b | 0.153 | marcksb | -0.017 |
| si:rp71-17i16.4 | 0.150 | si:dkey-276j7.1 | -0.017 |
| tnni2a.2 | 0.147 | tubb5 | -0.017 |
| ankrd24 | 0.144 | ywhaqb | -0.017 |
| pdlim3b | 0.144 | gng3 | -0.016 |
| mybpha | 0.138 | si:dkeyp-75h12.5 | -0.016 |
| smpx | 0.136 | stx1b | -0.016 |
| cav3 | 0.133 | celf2 | -0.015 |
| cavin4a | 0.133 | elavl4 | -0.015 |
| cavin1a | 0.131 | fabp3 | -0.015 |
| desma | 0.128 | mdka | -0.015 |
| CU681836.1 | 0.127 | myt1b | -0.015 |
| casq2 | 0.126 | nat8l | -0.015 |
| pvalb7 | 0.126 | si:ch211-137a8.4 | -0.015 |
| tnni2b.1 | 0.126 | tuba1c | -0.015 |
| zgc:86709 | 0.125 | stxbp1a | -0.015 |
| hspb11 | 0.124 | aplp1 | -0.014 |
| synpo2lb | 0.124 | atrx | -0.014 |
| tpm2 | 0.124 | calm2a | -0.014 |
| myoz3a | 0.122 | cbx3a | -0.014 |
| chrna1 | 0.121 | cd99l2 | -0.014 |