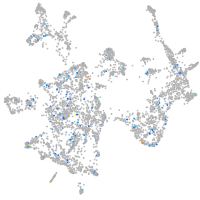
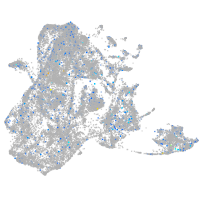
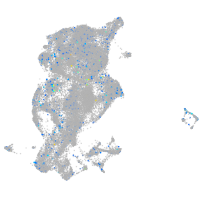
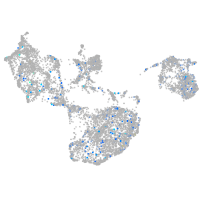
2-hydroxyacyl-CoA lyase 1
ZFIN 
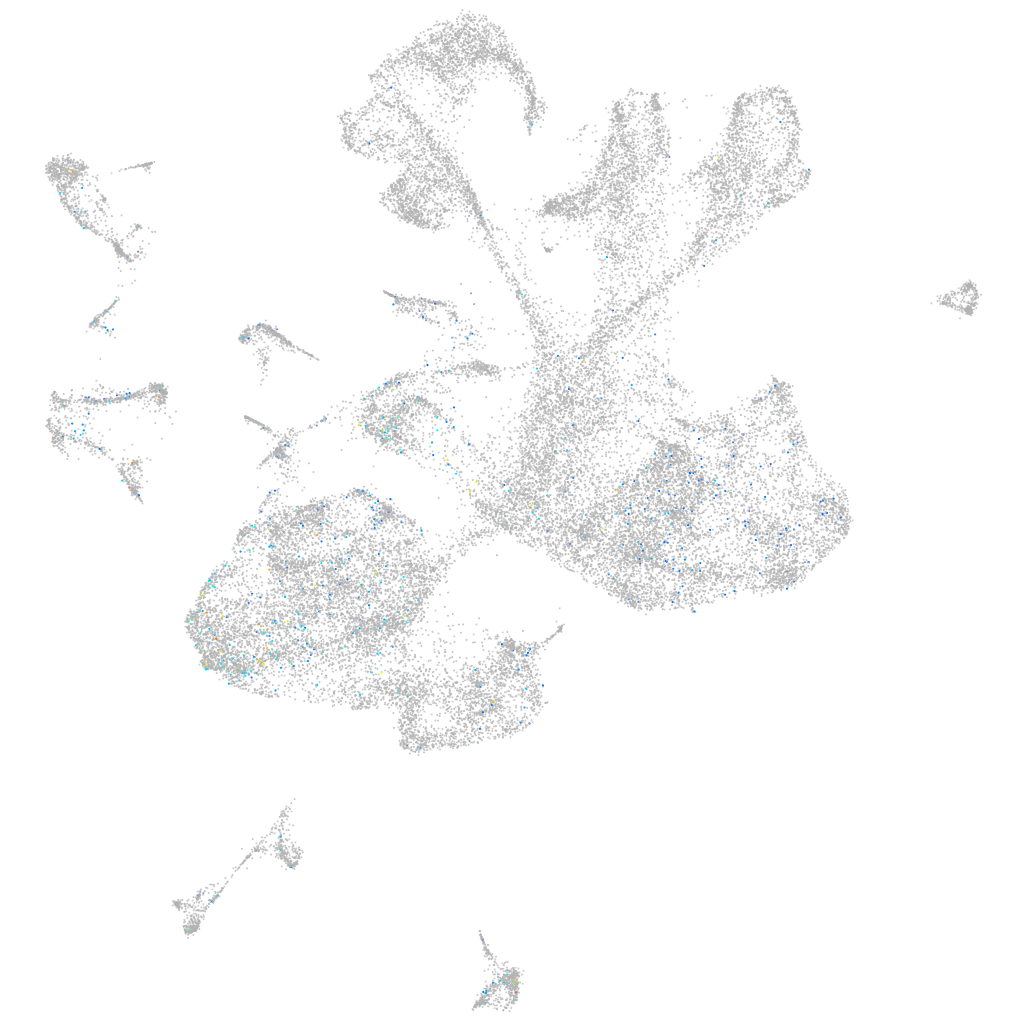
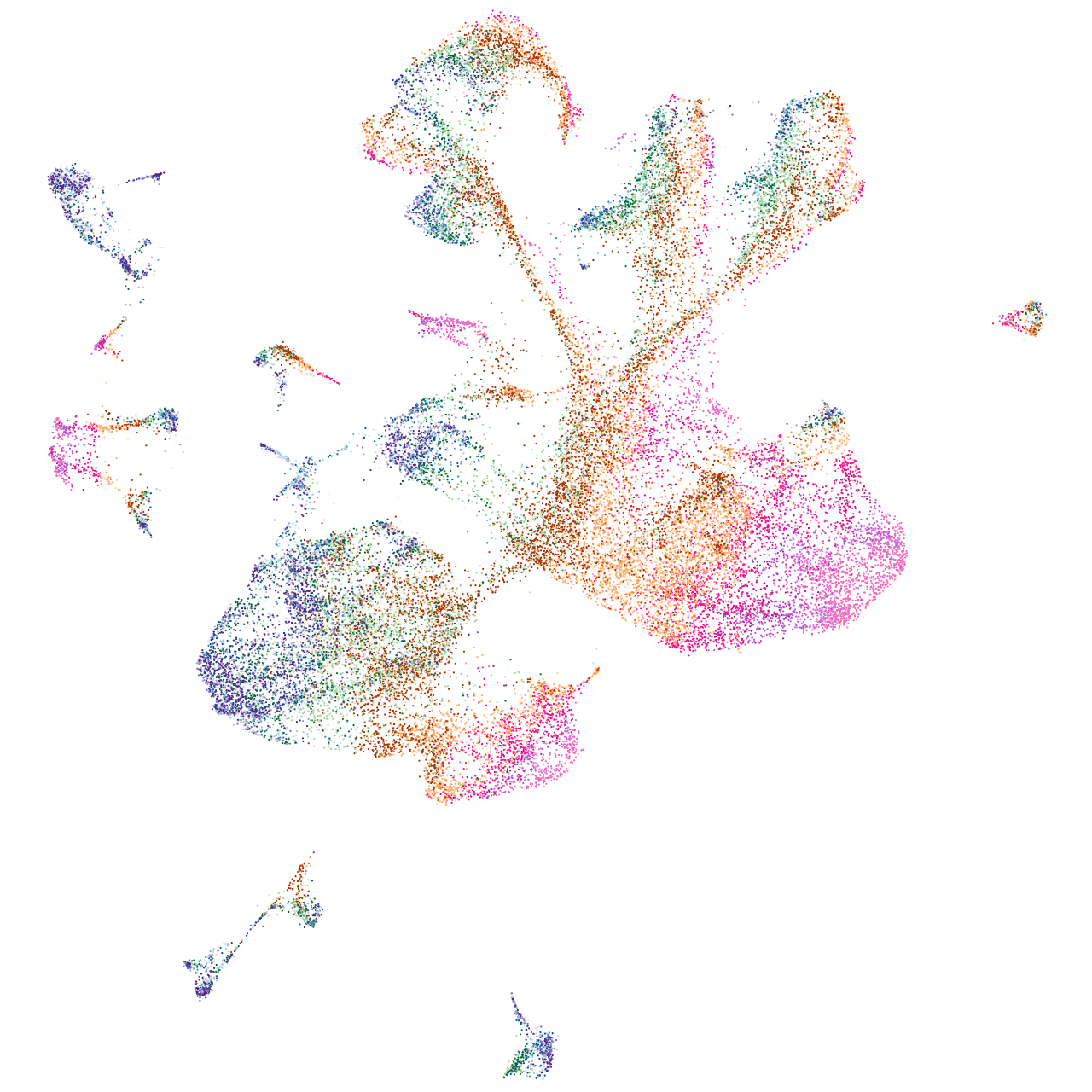
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| glula | 0.085 | ptmab | -0.064 |
| cx43 | 0.085 | elavl3 | -0.059 |
| hspb15 | 0.085 | si:ch211-222l21.1 | -0.056 |
| slc3a2a | 0.084 | h3f3d | -0.053 |
| slc6a11b | 0.082 | myt1b | -0.052 |
| ppap2d | 0.081 | tubb5 | -0.047 |
| mir301c | 0.080 | ptmaa | -0.045 |
| fabp7a | 0.077 | stxbp1a | -0.043 |
| slc4a4a | 0.076 | stx1b | -0.043 |
| gpr37l1b | 0.076 | cirbpb | -0.043 |
| si:ch211-66e2.5 | 0.076 | tubb2b | -0.043 |
| efhd1 | 0.076 | tmsb | -0.042 |
| mgll | 0.073 | gng3 | -0.042 |
| cdo1 | 0.073 | si:ch73-386h18.1 | -0.042 |
| cspg5b | 0.073 | snap25a | -0.041 |
| irge1 | 0.072 | hmgb3a | -0.040 |
| atp1a1b | 0.072 | stmn1b | -0.039 |
| sept8b | 0.072 | marcksb | -0.039 |
| slc1a2b | 0.071 | sncb | -0.038 |
| mfge8a | 0.071 | LOC100537384 | -0.038 |
| atp1b4 | 0.071 | chd4a | -0.038 |
| slc5a2 | 0.071 | elavl4 | -0.038 |
| cox4i2 | 0.070 | gng2 | -0.038 |
| si:ch1073-303k11.2 | 0.070 | onecut1 | -0.038 |
| hepacama | 0.069 | h3f3a | -0.038 |
| eno1b | 0.068 | ywhah | -0.037 |
| slc1a3b | 0.067 | zgc:100920 | -0.037 |
| pald1b | 0.067 | si:dkey-276j7.1 | -0.036 |
| gnai2a | 0.067 | rtn1a | -0.036 |
| ptgdsb.2 | 0.066 | onecut2 | -0.036 |
| cebpd | 0.065 | gap43 | -0.036 |
| BX004779.1 | 0.065 | hnrnpaba | -0.036 |
| s1pr1 | 0.064 | tmeff1b | -0.036 |
| slc38a3a | 0.064 | zc4h2 | -0.036 |
| cd63 | 0.064 | sox11b | -0.036 |