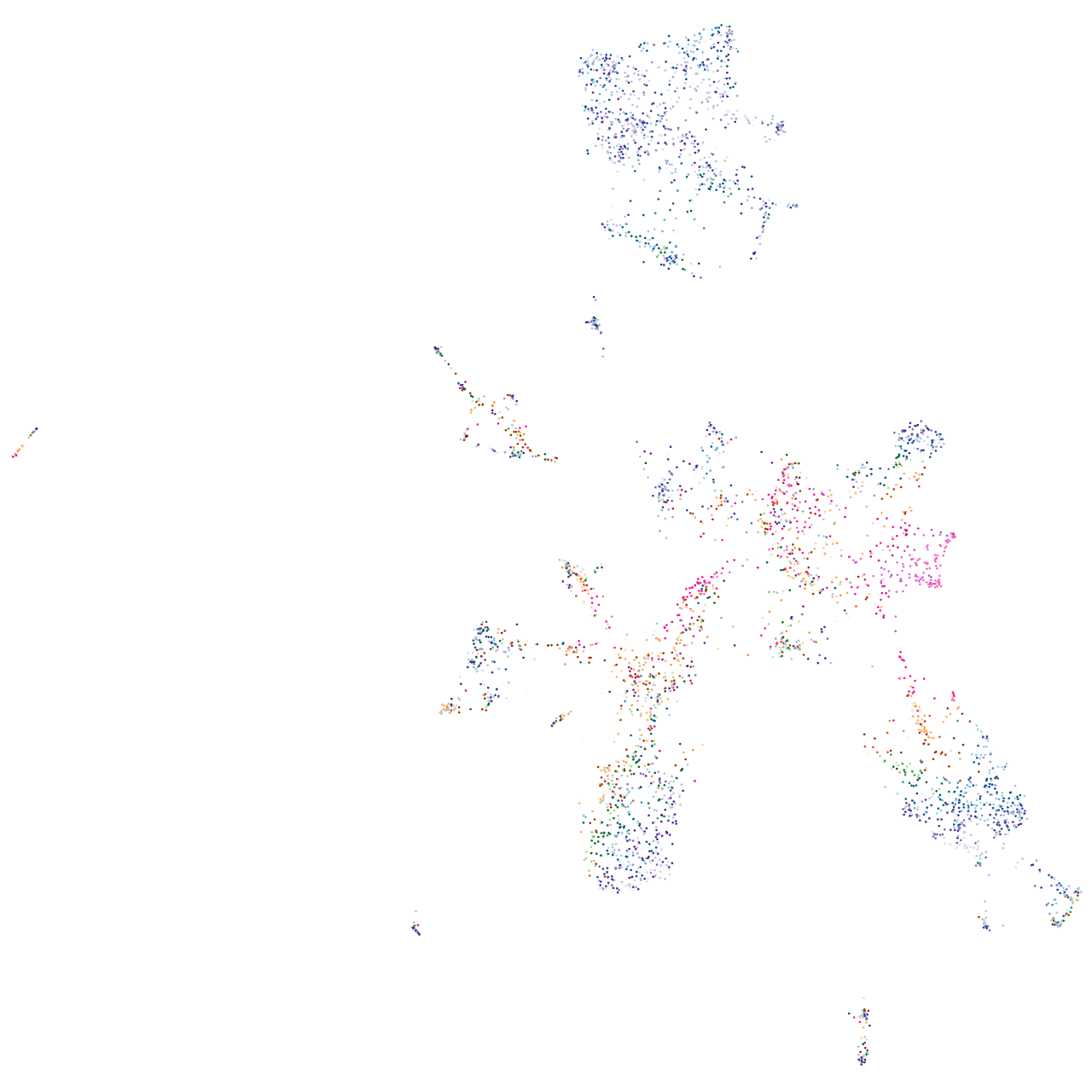
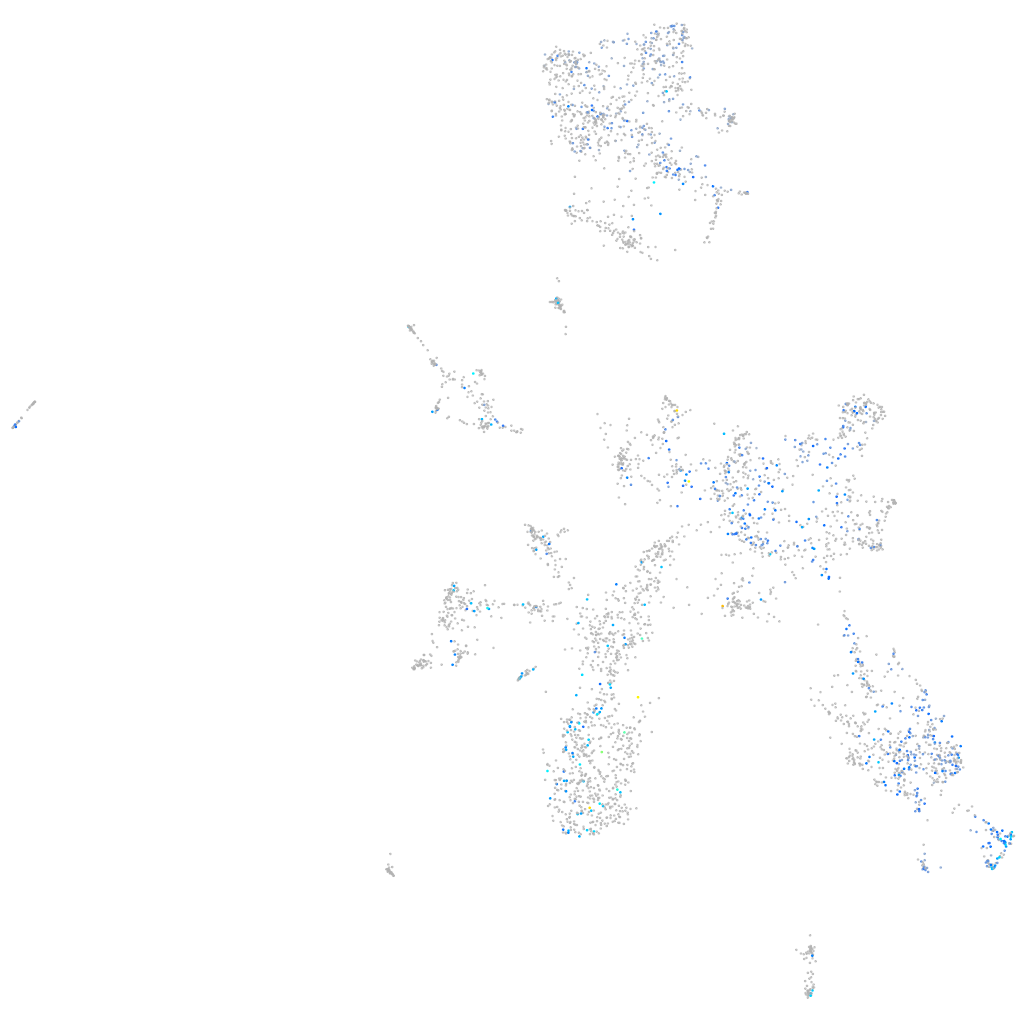
X-ray repair complementing defective repair in Chinese hamster cells 4
ZFIN 
Expression by stage/cluster


Correlated gene expression