transmembrane protein 42b
ZFIN 
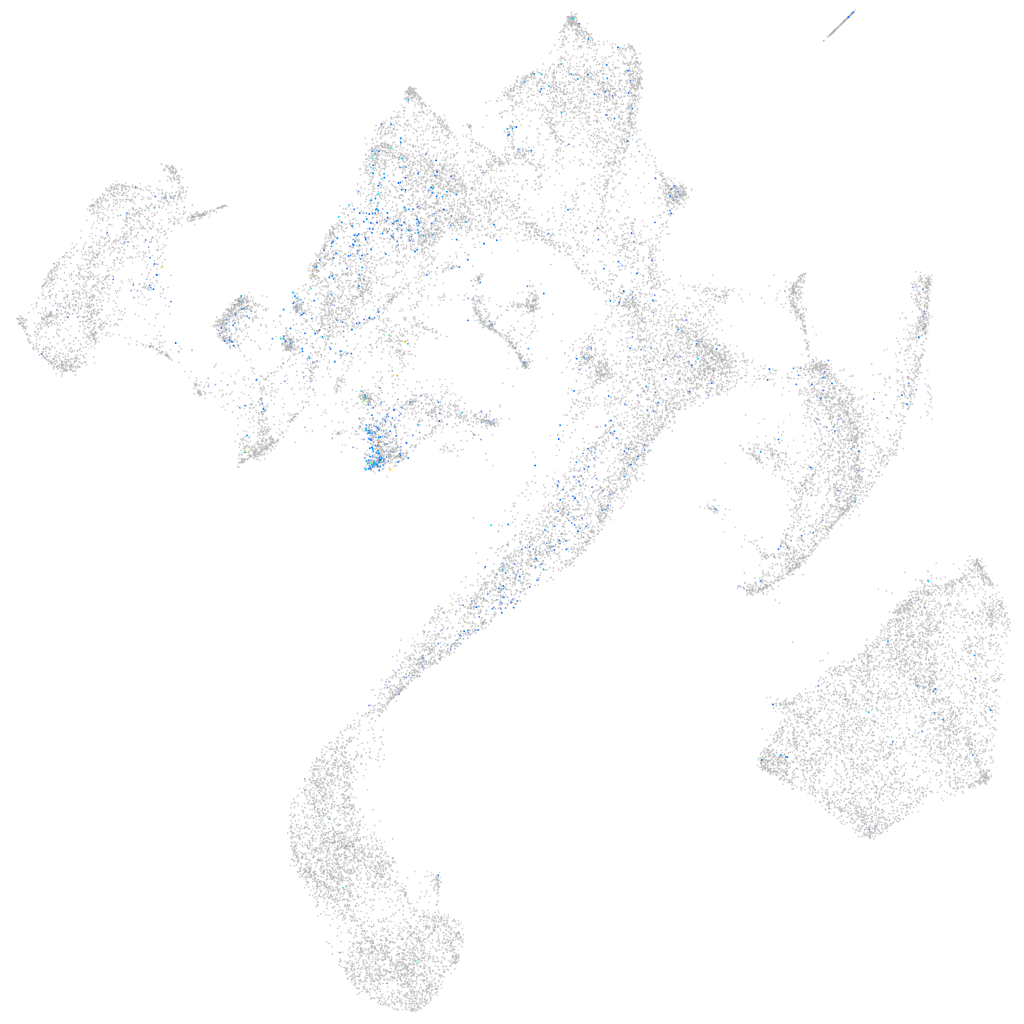
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| snap25a | 0.176 | apoeb | -0.067 |
| sncb | 0.159 | apoc1 | -0.064 |
| stxbp1a | 0.156 | cdx4 | -0.063 |
| sncgb | 0.154 | hspb1 | -0.060 |
| eno2 | 0.154 | BX927258.1 | -0.058 |
| rtn1b | 0.154 | NC-002333.4 | -0.057 |
| gng3 | 0.152 | tuba8l2 | -0.057 |
| vamp2 | 0.152 | cx43.4 | -0.055 |
| stmn2a | 0.149 | hes6 | -0.054 |
| elavl4 | 0.147 | zgc:110216 | -0.054 |
| stx1b | 0.146 | sp5l | -0.054 |
| stmn1b | 0.146 | asph | -0.053 |
| zgc:65894 | 0.145 | tbx16 | -0.053 |
| cdk5r2a | 0.143 | vox | -0.053 |
| nptna | 0.141 | hmgb2a | -0.052 |
| atp6v0cb | 0.140 | cdx1a | -0.052 |
| gabrg2 | 0.139 | crabp2b | -0.051 |
| syn2a | 0.139 | mki67 | -0.051 |
| vsnl1b | 0.139 | si:ch211-152c2.3 | -0.050 |
| snap25b | 0.137 | actn3a | -0.050 |
| cplx2 | 0.137 | si:ch211-255p10.3 | -0.050 |
| SRCIN1 | 0.135 | si:ch73-281n10.2 | -0.049 |
| gapdhs | 0.135 | XLOC-025819 | -0.049 |
| cplx2l | 0.134 | anp32e | -0.049 |
| bcl2b | 0.134 | ube2c | -0.049 |
| scg2b | 0.134 | XLOC-005350 | -0.049 |
| calm1b | 0.134 | ved | -0.048 |
| fxyd6 | 0.133 | hnrnpm | -0.048 |
| map1aa | 0.132 | tpx2 | -0.047 |
| evlb | 0.132 | zgc:110425 | -0.047 |
| syngr3a | 0.130 | CABZ01061524.1 | -0.047 |
| si:ch73-119p20.1 | 0.130 | msgn1 | -0.047 |
| gpm6aa | 0.129 | wu:fb97g03 | -0.046 |
| rnasekb | 0.129 | ccnb1 | -0.046 |
| ckbb | 0.128 | lig1 | -0.046 |