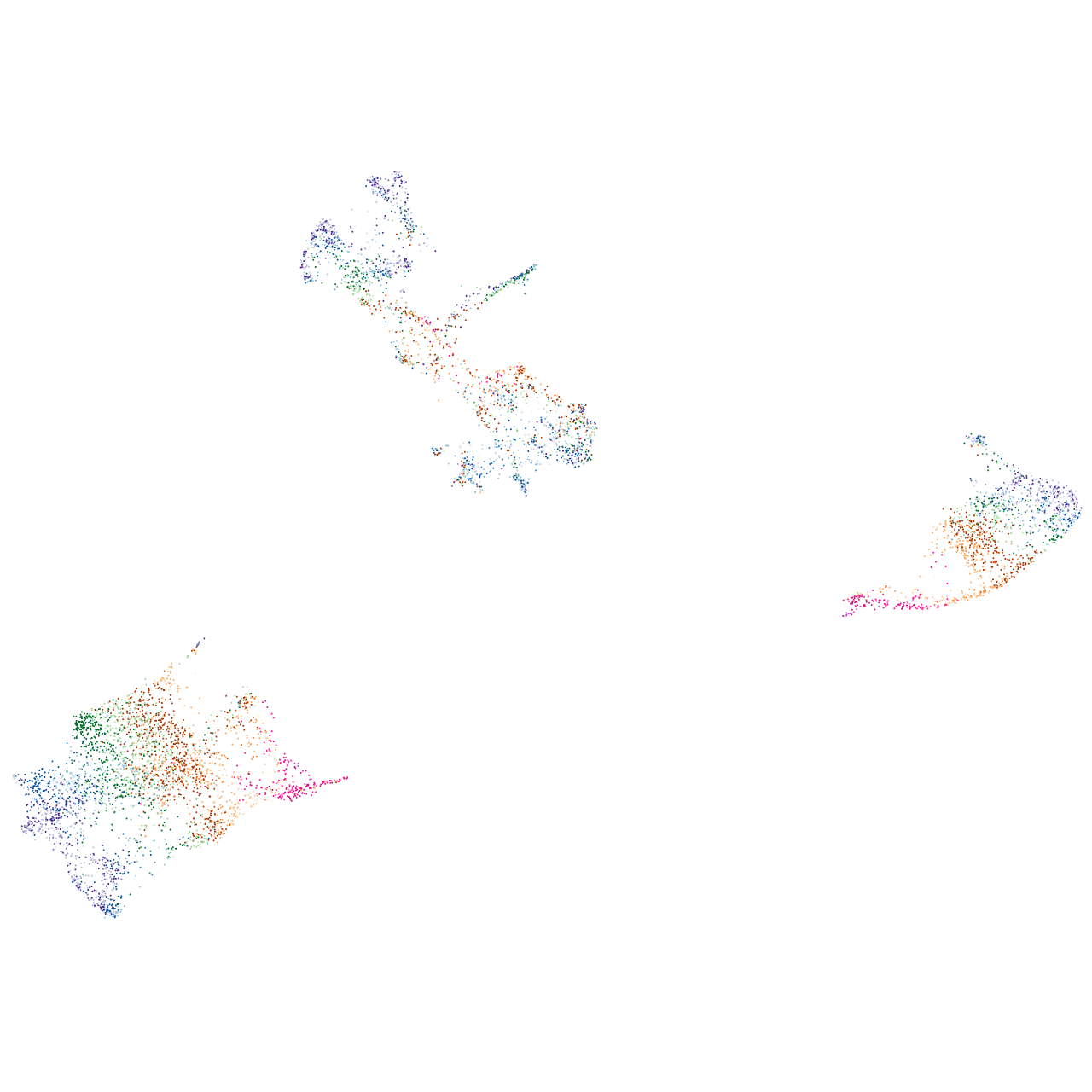
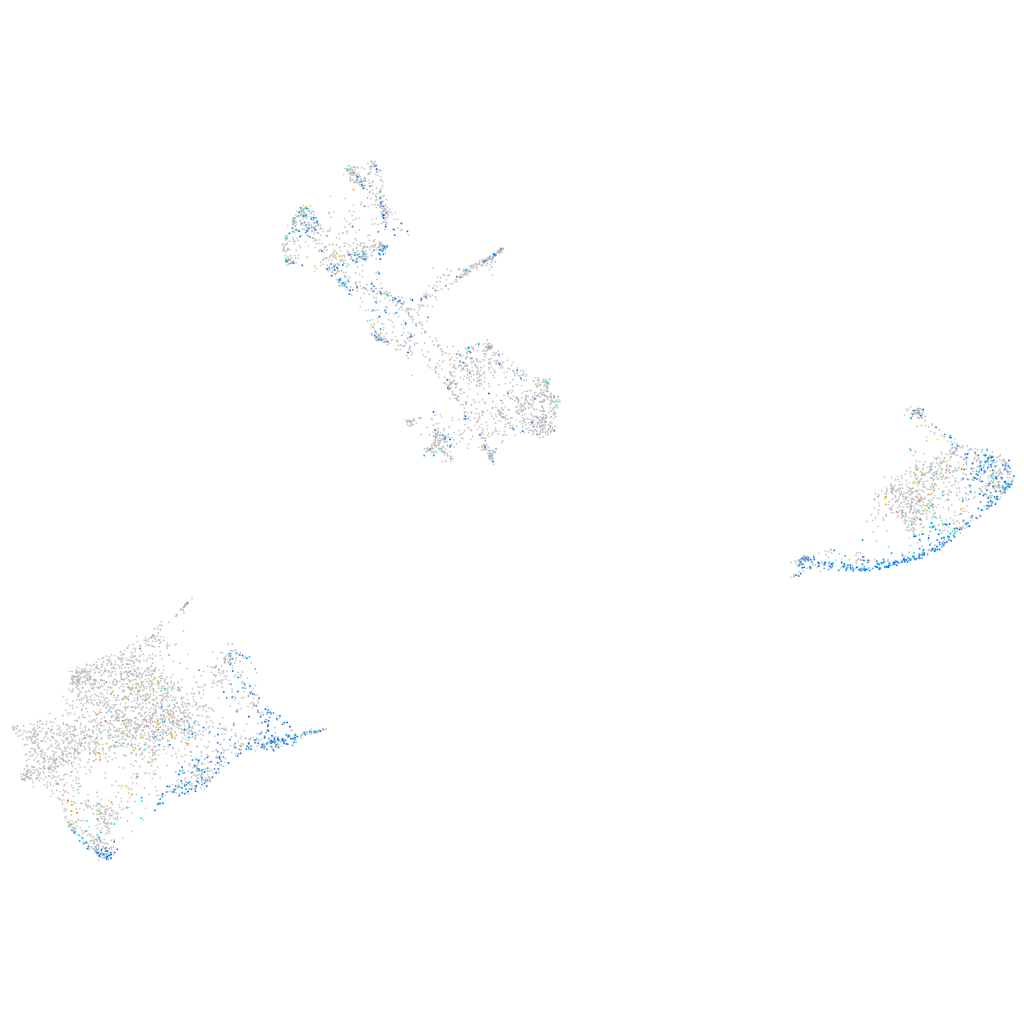
"solute carrier organic anion transporter family, member 1C1"
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| bace2 | 0.204 | ptmaa | -0.111 |
| SPAG9 | 0.195 | tuba1c | -0.098 |
| atp6ap1b | 0.192 | tmsb | -0.089 |
| trpm1b | 0.189 | nova2 | -0.089 |
| tspan36 | 0.185 | CABZ01021592.1 | -0.088 |
| fuca1.1 | 0.184 | pvalb2 | -0.088 |
| tyr | 0.182 | uraha | -0.087 |
| slc3a2a | 0.180 | pvalb1 | -0.084 |
| tmem243b | 0.179 | stmn1b | -0.084 |
| ctsba | 0.176 | si:ch211-222l21.1 | -0.082 |
| pttg1ipb | 0.175 | hbbe1.3 | -0.082 |
| lamp1a | 0.173 | hbae3 | -0.079 |
| slc45a2 | 0.172 | hbae1.1 | -0.078 |
| canx | 0.172 | hmgb3a | -0.076 |
| slc37a2 | 0.171 | si:ch73-1a9.3 | -0.076 |
| ctsla | 0.170 | gpm6aa | -0.075 |
| psap | 0.170 | elavl3 | -0.075 |
| syngr1a | 0.170 | si:dkey-251i10.2 | -0.075 |
| tfap2e | 0.169 | actc1b | -0.074 |
| mitfa | 0.168 | gng3 | -0.074 |
| atp11a | 0.168 | mylz3 | -0.072 |
| pcdh10a | 0.168 | hmgb1b | -0.072 |
| atp6v0ca | 0.167 | epb41a | -0.068 |
| pim1 | 0.167 | ckma | -0.067 |
| tspan10 | 0.166 | CU467822.1 | -0.067 |
| zgc:110239 | 0.165 | snap25a | -0.067 |
| rnaseka | 0.165 | hmgn2 | -0.067 |
| cdh1 | 0.163 | elavl4 | -0.066 |
| si:zfos-943e10.1 | 0.163 | krt4 | -0.065 |
| xbp1 | 0.160 | tnni2a.4 | -0.064 |
| pcdh9 | 0.160 | mylpfa | -0.063 |
| BX571715.1 | 0.159 | marcksl1a | -0.062 |
| msx1b | 0.159 | zc4h2 | -0.061 |
| degs1 | 0.159 | hbbe1.1 | -0.061 |
| klf6a | 0.158 | sncb | -0.060 |