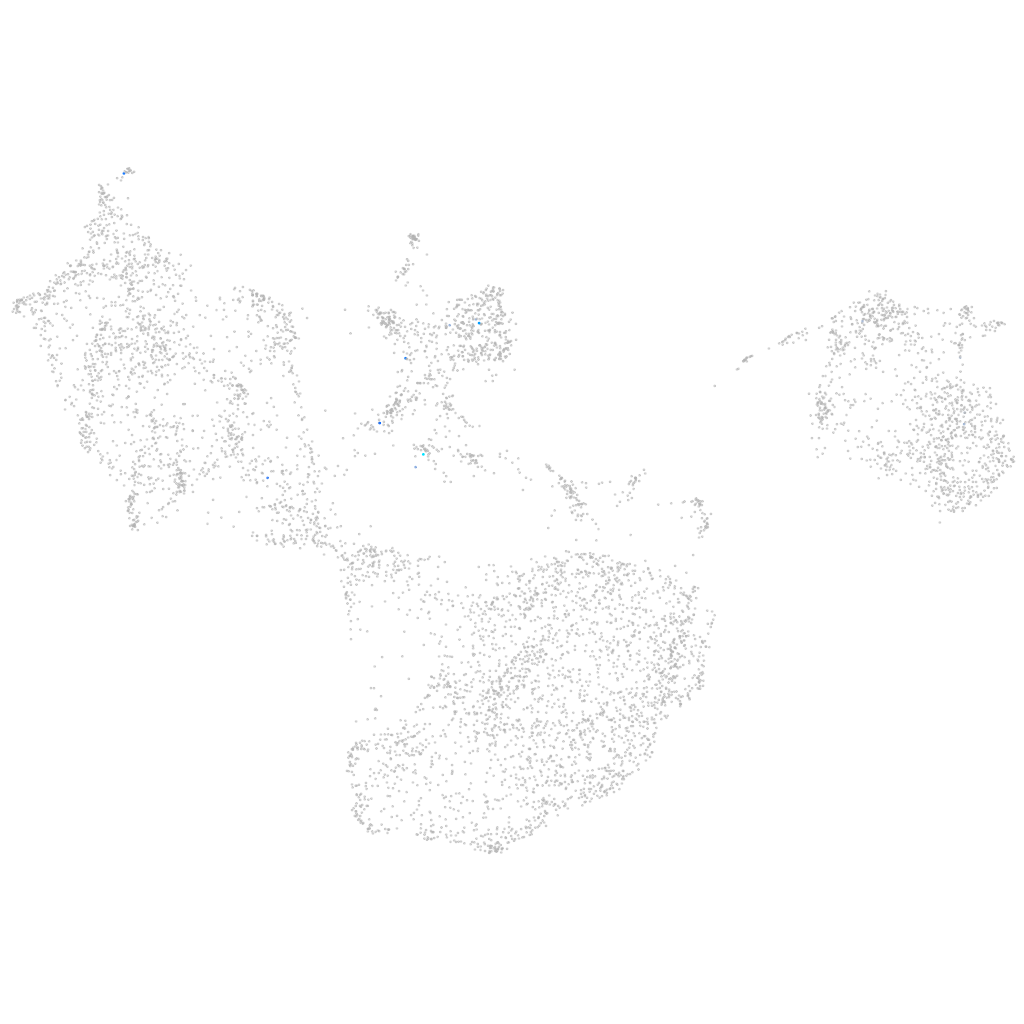
"notch receptor, like"
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| lyve1a | 0.644 | pfn1 | -0.037 |
| angpt2a | 0.632 | s100a10b | -0.036 |
| tnfaip2b | 0.495 | tmsb1 | -0.035 |
| kdr | 0.481 | cldni | -0.034 |
| acyp2 | 0.384 | cldn1 | -0.034 |
| gypc | 0.380 | cfl1l | -0.033 |
| myct1a | 0.376 | myh9a | -0.032 |
| rasip1 | 0.369 | epcam | -0.032 |
| plvapb | 0.354 | egfl6 | -0.032 |
| mfng | 0.334 | col18a1a | -0.032 |
| CT033796.1 | 0.331 | cyt1 | -0.032 |
| stab1 | 0.323 | thy1 | -0.032 |
| si:ch211-33e4.2 | 0.310 | cotl1 | -0.032 |
| clec14a | 0.297 | spaca4l | -0.031 |
| LOC101885919 | 0.296 | anxa1a | -0.031 |
| ecscr | 0.289 | apoeb | -0.031 |
| gpr182 | 0.287 | bcam | -0.031 |
| fgd5a | 0.283 | krt4 | -0.030 |
| pecam1 | 0.277 | arhgef5 | -0.030 |
| CABZ01053748.1 | 0.277 | si:dkey-102c8.3 | -0.029 |
| nxph4 | 0.274 | pak2a | -0.029 |
| mrc1a | 0.256 | zgc:158343 | -0.028 |
| cldn5b | 0.252 | tp63 | -0.028 |
| tie1 | 0.243 | arhgdig | -0.028 |
| XLOC-015089 | 0.235 | ecrg4b | -0.028 |
| si:ch211-119o8.4 | 0.230 | sh3d21 | -0.028 |
| zgc:165508 | 0.229 | pycard | -0.027 |
| LOC101882178 | 0.229 | vaspb | -0.027 |
| flt4 | 0.226 | aqp3a | -0.027 |
| CABZ01029422.1 | 0.225 | zgc:153284 | -0.027 |
| arfgef3 | 0.217 | krt5 | -0.027 |
| CU927890.1 | 0.215 | fmodb | -0.027 |
| dysf | 0.207 | zgc:92380 | -0.026 |
| smad4a | 0.205 | zgc:101810 | -0.026 |
| CR388417.1 | 0.205 | icn | -0.026 |




