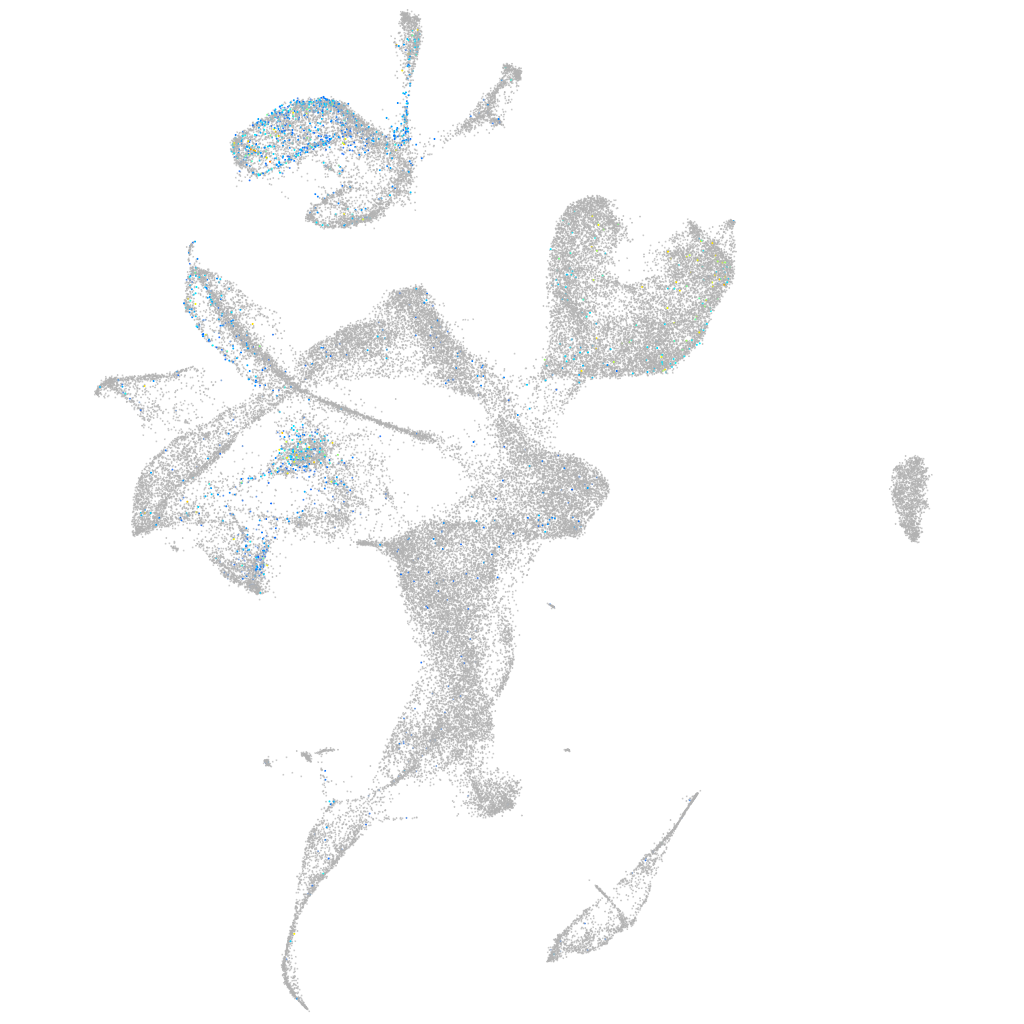
potassium channel tetramerization domain containing 4
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| ywhag2 | 0.164 | hmgb2a | -0.116 |
| zgc:65894 | 0.163 | pcna | -0.074 |
| tfap2a | 0.158 | stmn1a | -0.069 |
| pax10 | 0.157 | rrm1 | -0.068 |
| stxbp1a | 0.155 | ahcy | -0.068 |
| elavl3 | 0.154 | mki67 | -0.067 |
| gng3 | 0.151 | chaf1a | -0.067 |
| rtn1b | 0.150 | ccnd1 | -0.066 |
| slc32a1 | 0.149 | mcm7 | -0.066 |
| olfm2a | 0.147 | nutf2l | -0.066 |
| stmn2a | 0.146 | msi1 | -0.065 |
| scg2b | 0.143 | msna | -0.064 |
| syt1a | 0.143 | hmga1a | -0.064 |
| sncb | 0.141 | ccna2 | -0.064 |
| stx1b | 0.140 | dut | -0.063 |
| maptb | 0.140 | banf1 | -0.063 |
| elavl4 | 0.137 | lbr | -0.062 |
| slc35g2b | 0.136 | lig1 | -0.061 |
| stmn1b | 0.136 | dek | -0.060 |
| tfap2e | 0.134 | her15.1 | -0.060 |
| celf5a | 0.133 | mdka | -0.059 |
| slc6a1a | 0.132 | cks1b | -0.059 |
| zgc:153426 | 0.131 | selenoh | -0.059 |
| snap25a | 0.130 | tuba8l4 | -0.059 |
| gad2 | 0.128 | tuba8l | -0.058 |
| meis2b | 0.127 | rrm2 | -0.057 |
| tfap2b | 0.125 | zgc:110216 | -0.057 |
| fez1 | 0.124 | fen1 | -0.057 |
| c1ql1a | 0.124 | mibp | -0.057 |
| syn2a | 0.123 | mcm6 | -0.056 |
| meis2a | 0.123 | cdca7b | -0.056 |
| sprn | 0.123 | nasp | -0.056 |
| ppp1r14ba | 0.122 | mcm2 | -0.056 |
| gabrb4 | 0.122 | id1 | -0.055 |
| eno2 | 0.122 | dlgap5 | -0.055 |