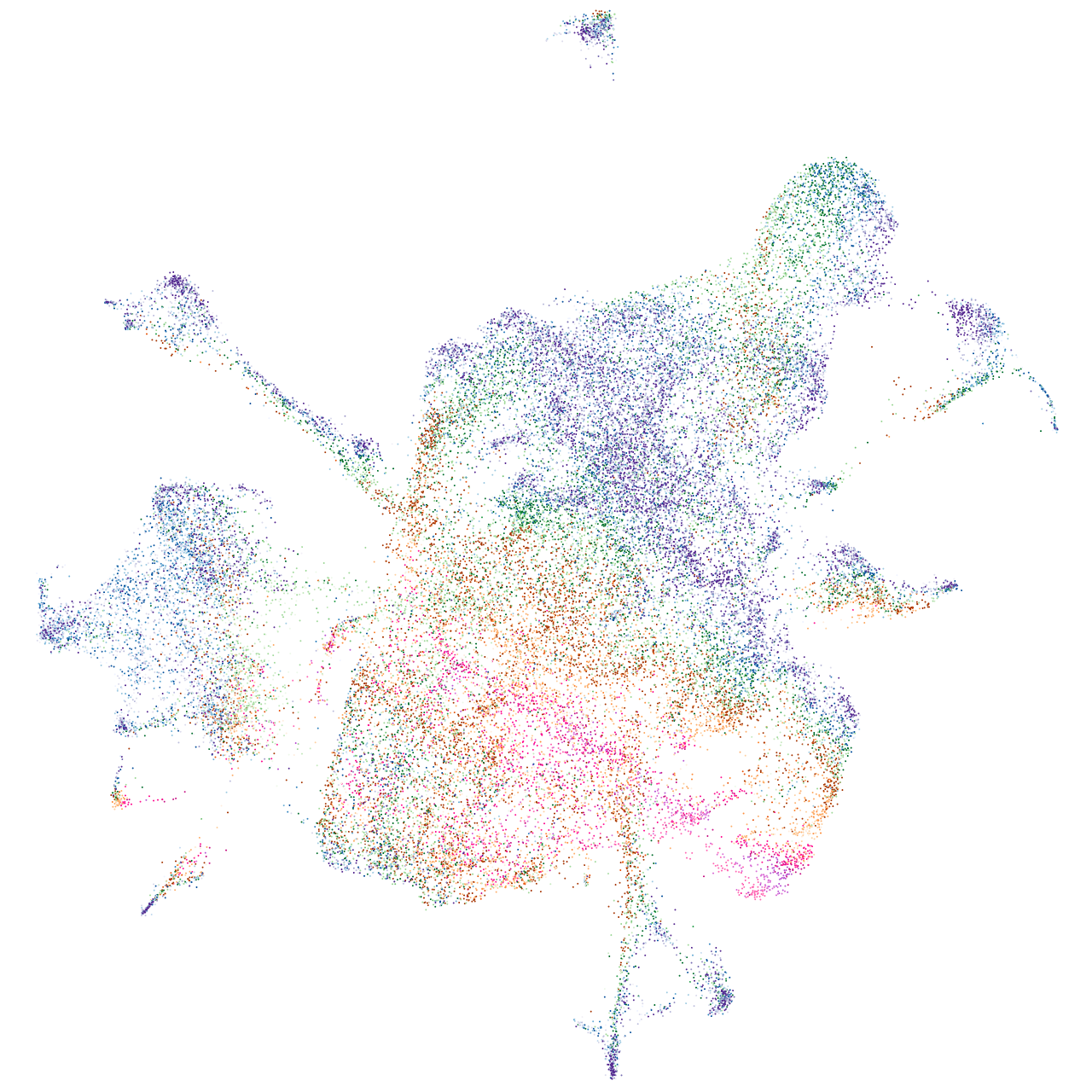
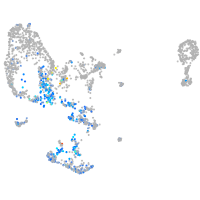
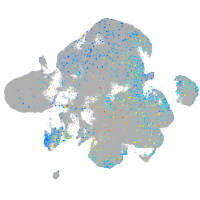
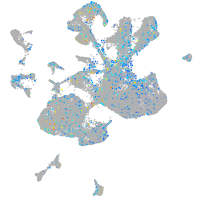
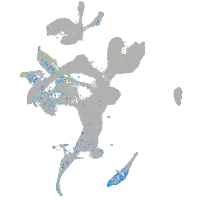
erythrocyte membrane protein band 4.1 like 4A
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| elavl3 | 0.088 | pmp22a | -0.041 |
| CU855936.2 | 0.085 | fibina | -0.036 |
| si:ch211-216p19.5 | 0.084 | CU929237.1 | -0.035 |
| podxl | 0.080 | LO018188.1 | -0.034 |
| gpm6aa | 0.074 | foxc1b | -0.034 |
| zgc:158291 | 0.074 | fgfbp2b | -0.032 |
| scrt2 | 0.074 | col9a1a | -0.030 |
| fez1 | 0.074 | rarga | -0.030 |
| fat2 | 0.074 | six1a | -0.030 |
| tuba1c | 0.072 | rpl36 | -0.029 |
| LOC103909274 | 0.070 | cnmd | -0.028 |
| tuba1a | 0.069 | rplp1 | -0.028 |
| tmsb | 0.069 | col9a3 | -0.027 |
| isl2b | 0.068 | ecrg4a | -0.026 |
| tmem88b | 0.067 | sox9a | -0.026 |
| mov10b.1 | 0.067 | col2a1a | -0.026 |
| stmn1b | 0.067 | VIT | -0.026 |
| hoxc11a | 0.067 | rpl26 | -0.025 |
| bcam | 0.067 | zgc:114188 | -0.025 |
| jam2b | 0.066 | col9a2 | -0.025 |
| si:dkey-56m19.5 | 0.066 | rpl23a | -0.024 |
| gng3 | 0.065 | loxl1 | -0.024 |
| nova2 | 0.064 | rps15 | -0.024 |
| alcamb | 0.064 | phlda3 | -0.024 |
| inab | 0.064 | hapln1a | -0.024 |
| nsg2 | 0.064 | si:dkey-151g10.6 | -0.023 |
| rtn1a | 0.063 | rplp2 | -0.023 |
| marcksl1b | 0.062 | col2a1b | -0.023 |
| mllt11 | 0.061 | rpl7 | -0.022 |
| elavl4 | 0.061 | rpl27a | -0.022 |
| cavin2b | 0.061 | rpl18 | -0.022 |
| lrrc74a | 0.061 | rargb | -0.022 |
| islr2 | 0.061 | barx1 | -0.022 |
| si:dkey-280e21.3 | 0.060 | gsc | -0.022 |
| rnasekb | 0.060 | aspn | -0.022 |