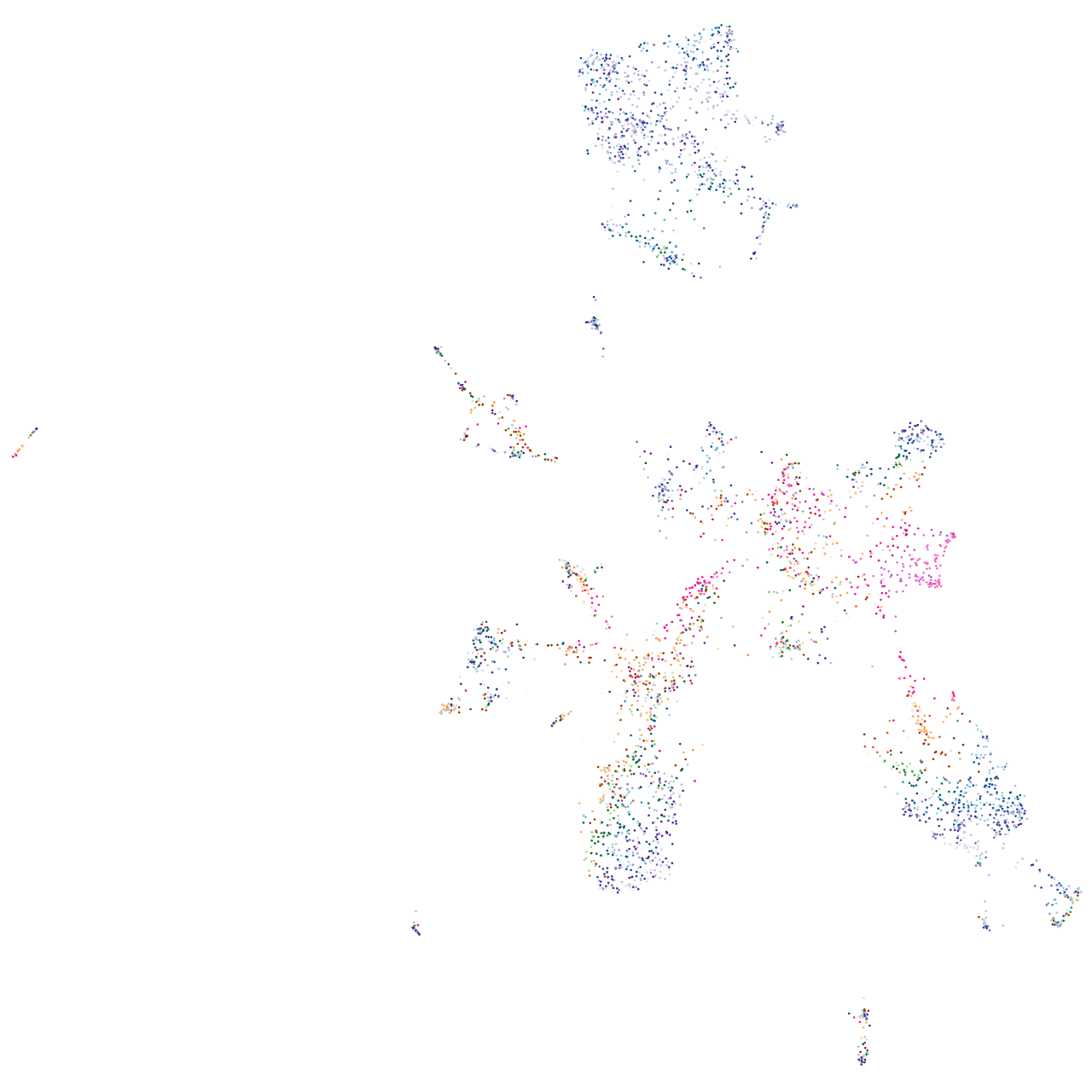
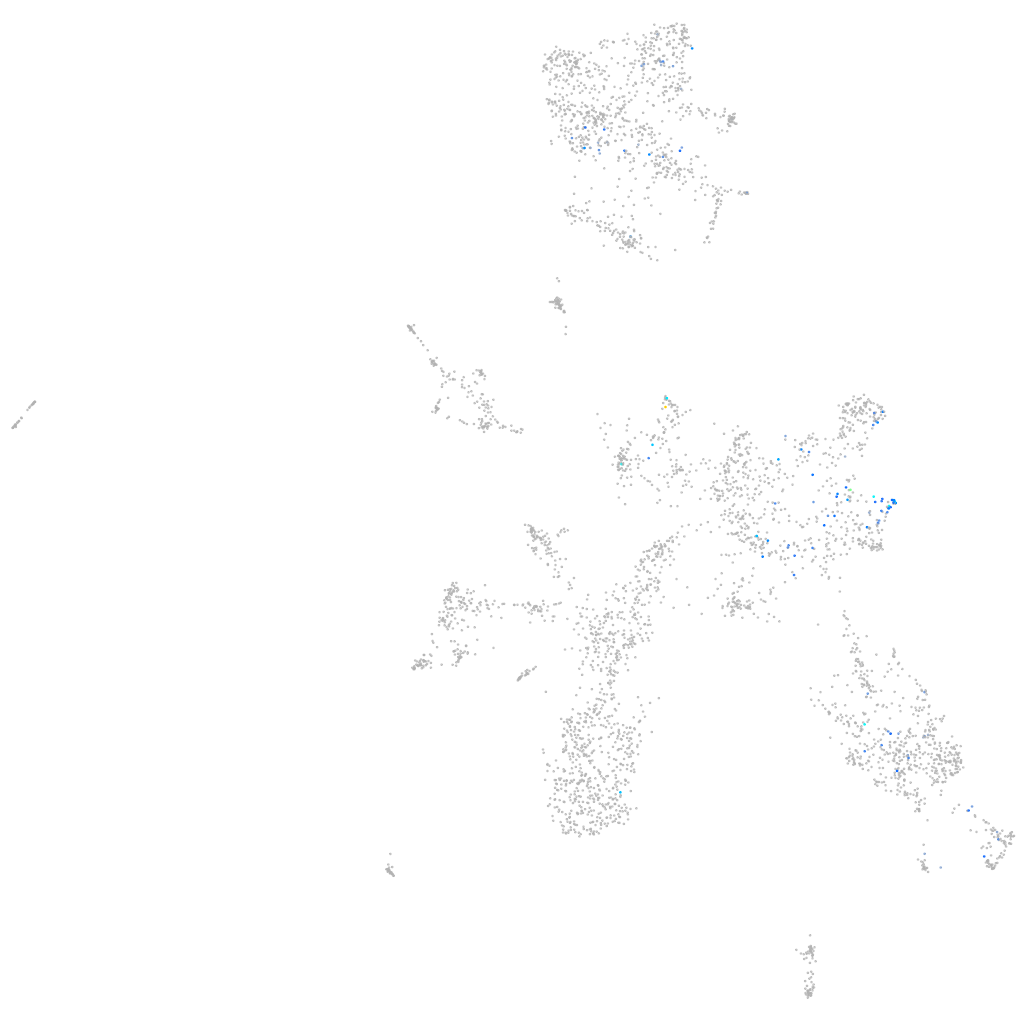
distal-less homeobox 4a
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| CU302323.1 | 0.283 | cldnh | -0.137 |
| nradd | 0.281 | rnasekb | -0.102 |
| CABZ01079192.1 | 0.281 | rtn1a | -0.098 |
| ftr76 | 0.279 | txn | -0.092 |
| adgre5a | 0.271 | insm1a | -0.091 |
| XLOC-043778 | 0.254 | fxyd1 | -0.090 |
| msx1b | 0.252 | gapdhs | -0.086 |
| tfap2a | 0.252 | vamp2 | -0.084 |
| asb11 | 0.241 | gnb1a | -0.082 |
| thy1 | 0.238 | tuba1c | -0.081 |
| igfn1.4 | 0.233 | elavl3 | -0.080 |
| fgfbp2a | 0.228 | tspan2a | -0.080 |
| klf17 | 0.215 | atp6v0cb | -0.078 |
| zgc:165539 | 0.212 | insm1b | -0.078 |
| si:busm1-160c18.6 | 0.210 | atp6v1e1b | -0.077 |
| tacr3a | 0.209 | scg2b | -0.076 |
| FP103005.1 | 0.208 | zgc:65894 | -0.075 |
| tpm4a | 0.206 | LOC100537342 | -0.075 |
| tfap2c | 0.202 | tuba1a | -0.075 |
| ptgdsb.1 | 0.202 | anxa5b | -0.074 |
| sema3h | 0.195 | eno1a | -0.073 |
| BX510337.1 | 0.195 | gng3 | -0.073 |
| pax7b | 0.192 | flrt1b | -0.073 |
| gas1a | 0.186 | chga | -0.072 |
| cx44.2 | 0.186 | reep5 | -0.072 |
| gyg2 | 0.185 | stmn1b | -0.072 |
| dlx3b | 0.184 | calm2b | -0.071 |
| si:ch211-73m21.1 | 0.181 | clstn1 | -0.071 |
| si:dkey-233e3.3 | 0.180 | tmsb | -0.070 |
| ptgs2a | 0.177 | ptmaa | -0.070 |
| nid2a | 0.177 | acbd7 | -0.069 |
| KCTD15 | 0.172 | si:dkeyp-75h12.5 | -0.069 |
| pmp22a | 0.171 | fstl5 | -0.069 |
| pgfb | 0.171 | scg2a | -0.069 |
| yap1 | 0.169 | nfe2l2a | -0.069 |