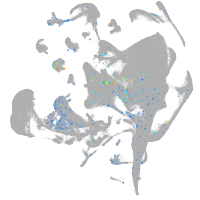
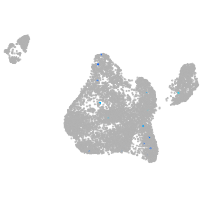
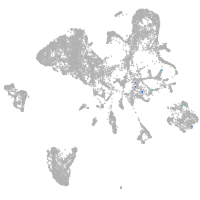
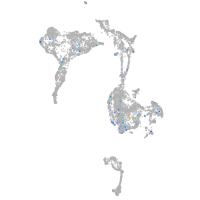
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| slc4a4a | 0.125 | ptmab | -0.095 |
| gpr37l1b | 0.125 | si:ch211-222l21.1 | -0.088 |
| FP102888.1 | 0.120 | h3f3a | -0.080 |
| cx43 | 0.120 | cirbpb | -0.073 |
| pbxip1b | 0.111 | tubb2b | -0.072 |
| slc3a2a | 0.110 | hnrnpaba | -0.070 |
| si:ch211-180a12.2 | 0.109 | hsp90ab1 | -0.070 |
| slc6a11b | 0.109 | hnrnpa0l | -0.070 |
| hepacama | 0.109 | h3f3d | -0.069 |
| atp1a1b | 0.109 | h2afvb | -0.066 |
| pygb | 0.107 | khdrbs1a | -0.062 |
| si:ch211-66e2.5 | 0.106 | rps20 | -0.062 |
| si:dkey-245n4.2 | 0.106 | hnrnpa0b | -0.062 |
| si:ch1073-303k11.2 | 0.106 | fabp3 | -0.060 |
| slc1a2b | 0.104 | faua | -0.060 |
| slc10a1 | 0.103 | rpl24 | -0.059 |
| atp1b4 | 0.102 | elavl3 | -0.059 |
| zgc:153704 | 0.102 | marcksb | -0.058 |
| ptn | 0.100 | ptmaa | -0.058 |
| cldn7a | 0.100 | hmgn2 | -0.057 |
| ptgdsb.2 | 0.098 | rpl19 | -0.057 |
| cav1 | 0.097 | rps21 | -0.057 |
| ppap2d | 0.097 | cirbpa | -0.056 |
| eno1b | 0.097 | rpl21 | -0.056 |
| hspb15 | 0.096 | rps23 | -0.055 |
| cox4i2 | 0.094 | si:ch211-288g17.3 | -0.055 |
| sept8b | 0.091 | rpl22 | -0.055 |
| glula | 0.090 | rps12 | -0.055 |
| tmem145 | 0.089 | hmga1a | -0.054 |
| lpin1 | 0.089 | sumo3a | -0.054 |
| slc1a3b | 0.089 | rpl39 | -0.054 |
| pald1b | 0.089 | snrpf | -0.053 |
| cyp2ad3 | 0.089 | rps27.1 | -0.053 |
| endouc | 0.089 | hmgb2b | -0.052 |
| fabp7a | 0.088 | ddx39ab | -0.052 |