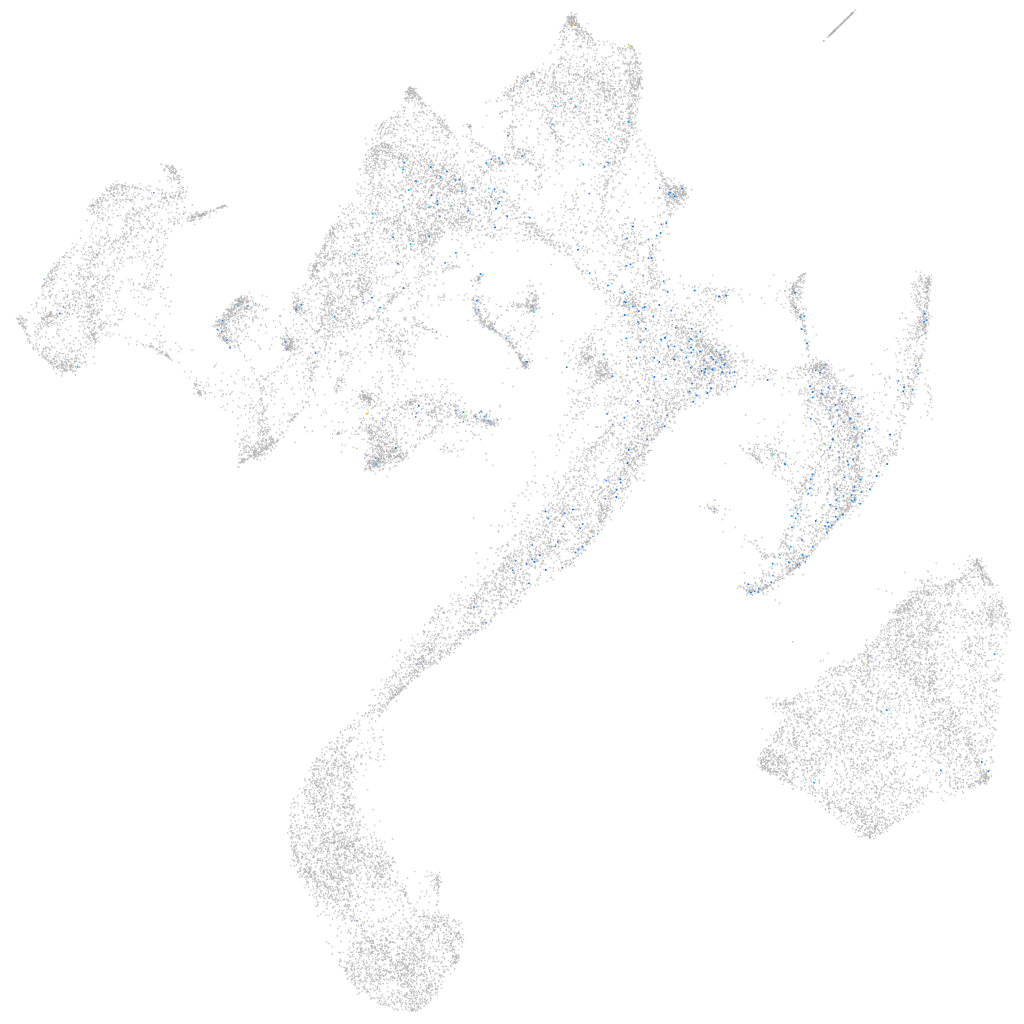
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| hnrnpa0l | 0.089 | ckmb | -0.055 |
| h3f3a | 0.083 | ckma | -0.055 |
| zgc:56493 | 0.078 | gapdh | -0.054 |
| lmx1al | 0.078 | atp2a1 | -0.053 |
| dut | 0.076 | ak1 | -0.053 |
| eif3ba | 0.071 | tnnc2 | -0.052 |
| zgc:103625 | 0.070 | neb | -0.051 |
| fn1b | 0.069 | actc1b | -0.051 |
| dram1 | 0.068 | ldb3b | -0.050 |
| ewsr1b | 0.068 | si:ch73-367p23.2 | -0.050 |
| myo15b | 0.068 | aldoab | -0.049 |
| fshb | 0.067 | mylpfa | -0.049 |
| si:ch1073-429i10.3.1 | 0.067 | actn3a | -0.049 |
| naca | 0.067 | smyd1a | -0.048 |
| ran | 0.067 | actn3b | -0.048 |
| tma16 | 0.066 | myom1a | -0.048 |
| eif3m | 0.066 | mylz3 | -0.048 |
| her1 | 0.065 | tnnt3a | -0.048 |
| rrm2 | 0.065 | CABZ01078594.1 | -0.047 |
| rpl10a | 0.064 | tmem38a | -0.047 |
| tmem30c | 0.064 | cav3 | -0.047 |
| eif3s6ip | 0.064 | si:ch211-266g18.10 | -0.047 |
| hsp90ab1 | 0.063 | pgam2 | -0.047 |
| pin1 | 0.063 | pvalb2 | -0.047 |
| tubb4b | 0.063 | tmod4 | -0.046 |
| tspan7 | 0.063 | glud1b | -0.046 |
| rps7 | 0.062 | pvalb1 | -0.046 |
| rpl12 | 0.062 | tnnt3b | -0.046 |
| CR318603.2 | 0.062 | XLOC-001975 | -0.045 |
| ubc | 0.062 | cavin4b | -0.045 |
| sall4 | 0.062 | casq2 | -0.045 |
| eef2b | 0.061 | acta1b | -0.045 |
| rps14 | 0.061 | eno1a | -0.045 |
| syngap1a | 0.061 | prx | -0.045 |
| hoxb10a | 0.061 | eno3 | -0.044 |