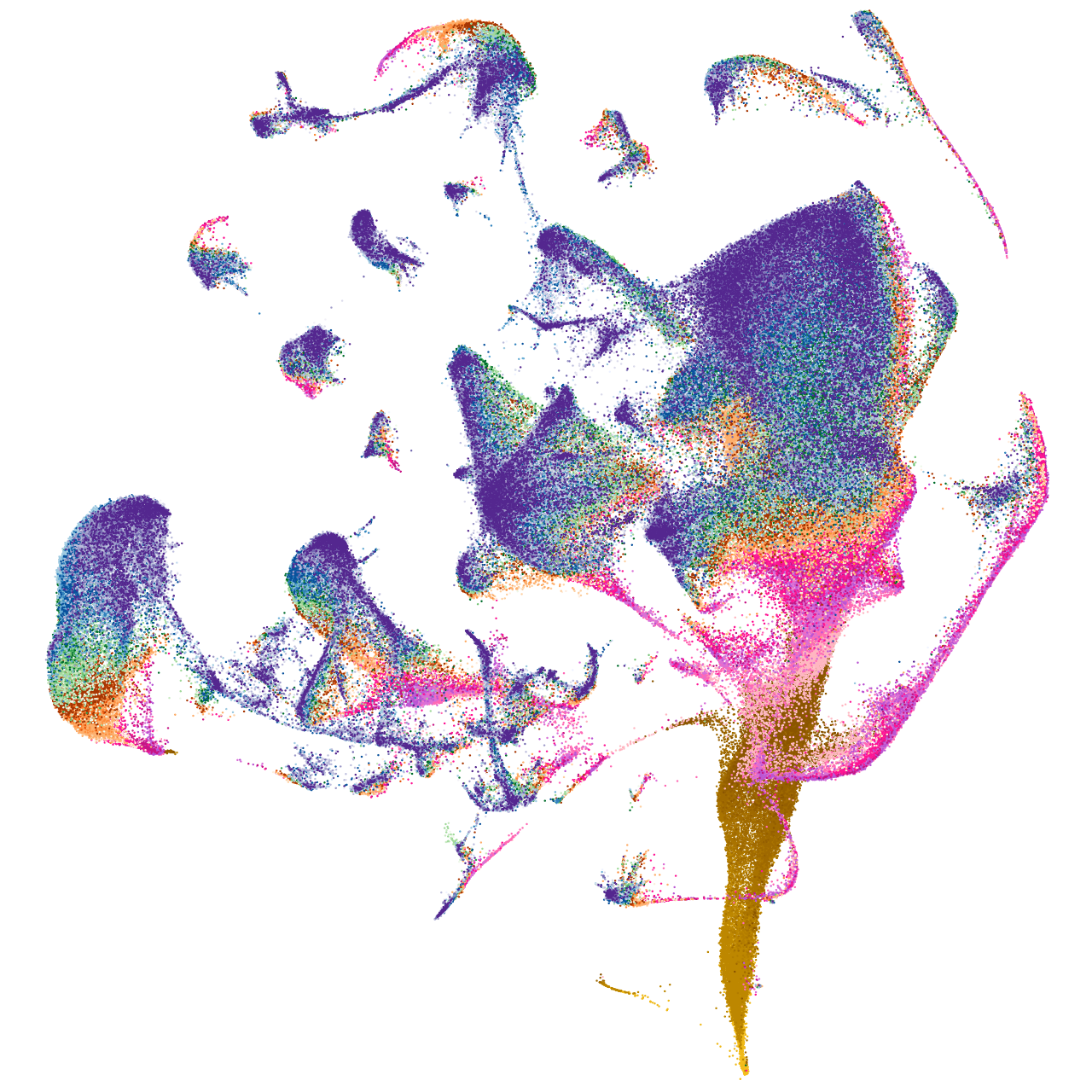WD repeat domain 78
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| daw1 | 0.245 | stmn1b | -0.021 |
| tbata | 0.235 | celf2 | -0.020 |
| morn3 | 0.228 | gpm6ab | -0.020 |
| foxj1a | 0.227 | rtn1b | -0.018 |
| enkur | 0.225 | ywhag2 | -0.018 |
| spata18 | 0.221 | atp1b2a | -0.017 |
| ccdc151 | 0.220 | elavl4 | -0.017 |
| si:ch211-155m12.5 | 0.217 | gng3 | -0.017 |
| si:dkey-27p23.3 | 0.214 | sypb | -0.017 |
| cfap126 | 0.213 | atp1a3a | -0.016 |
| ttc25 | 0.211 | mab21l1 | -0.016 |
| pacrg | 0.208 | ndrg4 | -0.016 |
| si:ch211-163l21.7 | 0.207 | snap25b | -0.016 |
| tekt1 | 0.203 | tmem59l | -0.016 |
| dnali1 | 0.202 | cyt1 | -0.015 |
| spag6 | 0.202 | elavl3 | -0.015 |
| tekt4 | 0.200 | krtt1c19e | -0.015 |
| ccdc114 | 0.199 | rbfox1 | -0.015 |
| si:dkey-148f10.4 | 0.199 | rtn1a | -0.015 |
| si:ch211-226m7.4 | 0.196 | stx1b | -0.015 |
| ccdc173 | 0.195 | ywhah | -0.015 |
| smkr1 | 0.194 | atp6v0cb | -0.014 |
| CFAP77 | 0.191 | camk2d2 | -0.014 |
| ak7a | 0.190 | cdk5r1b | -0.014 |
| efhc2 | 0.189 | col1a2 | -0.014 |
| cfap157 | 0.186 | gap43 | -0.014 |
| ccdc169 | 0.184 | mab21l2 | -0.014 |
| nme5 | 0.184 | mllt11 | -0.014 |
| ak8 | 0.183 | myt1b | -0.014 |
| ccdc65 | 0.180 | scrt2 | -0.014 |
| dnaaf1 | 0.180 | si:dkey-276j7.1 | -0.014 |
| ropn1l | 0.180 | sncgb | -0.014 |
| mycbpap | 0.179 | stmn2a | -0.014 |
| spaca9 | 0.179 | tubb5 | -0.014 |
| ak9 | 0.179 | zgc:153426 | -0.014 |