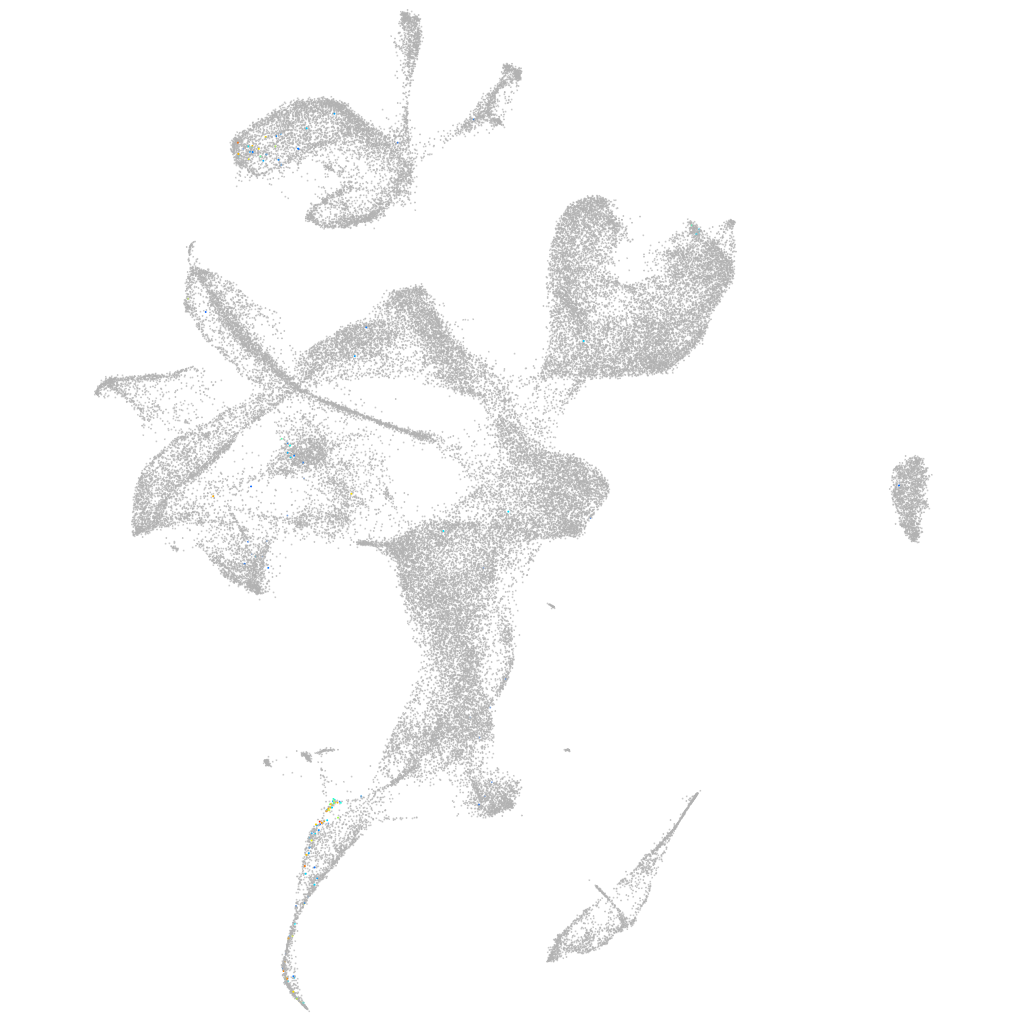
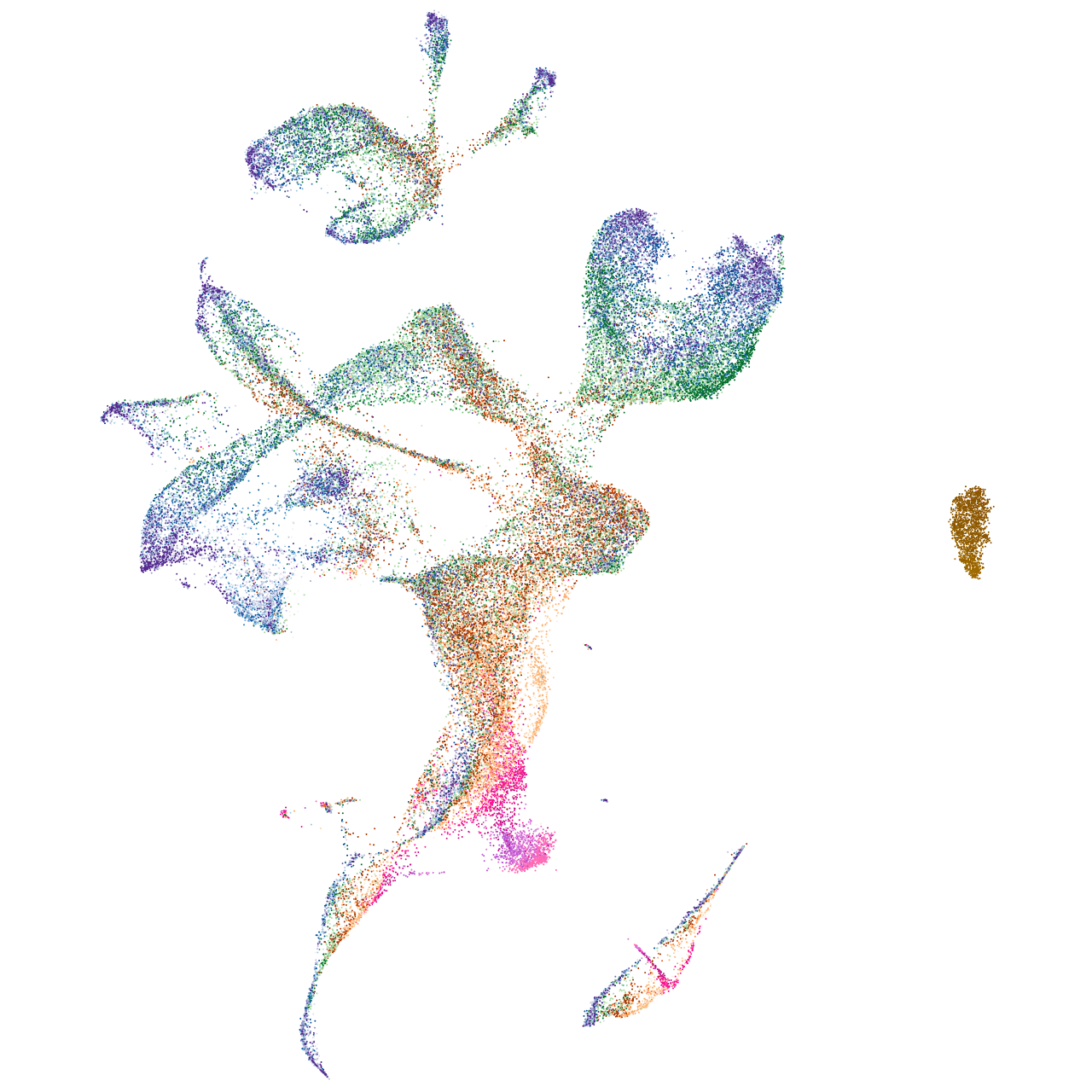thyrotropin-releasing hormone
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| c1qtnf1 | 0.264 | ptmab | -0.058 |
| thbs3b | 0.228 | si:ch73-1a9.3 | -0.057 |
| cldn7a | 0.182 | hmga1a | -0.040 |
| cldn19 | 0.182 | hmgn2 | -0.039 |
| artnb | 0.175 | h2afva | -0.037 |
| ccka | 0.171 | fabp7a | -0.037 |
| cyp24a1 | 0.157 | hnrnpaba | -0.037 |
| ldlrap1a | 0.156 | hmgb2a | -0.036 |
| oca2 | 0.152 | hnrnpa0b | -0.036 |
| tyrp1b | 0.152 | khdrbs1a | -0.036 |
| rab27a | 0.151 | hmgn6 | -0.036 |
| pmela | 0.151 | snrpf | -0.035 |
| dct | 0.148 | snrpd1 | -0.032 |
| LOC108191636 | 0.147 | hmgb2b | -0.032 |
| lmod1b | 0.147 | fabp3 | -0.032 |
| tspan36 | 0.144 | snrpe | -0.031 |
| fmn1 | 0.144 | h2afvb | -0.031 |
| srgn | 0.143 | si:ch211-288g17.3 | -0.031 |
| myocd | 0.139 | cirbpb | -0.030 |
| sytl2a | 0.138 | si:ch211-137a8.4 | -0.030 |
| cnn1b | 0.135 | si:ch73-281n10.2 | -0.028 |
| acta2 | 0.135 | hdac1 | -0.028 |
| bace2 | 0.131 | anp32e | -0.028 |
| myl9a | 0.131 | snrpb | -0.028 |
| gpr143 | 0.126 | cct4 | -0.027 |
| ugt1b5 | 0.124 | h3f3d | -0.027 |
| sytl2b | 0.123 | snrpg | -0.027 |
| agtrap | 0.122 | seta | -0.027 |
| tyrp1a | 0.122 | ran | -0.027 |
| rab38 | 0.121 | marcksb | -0.026 |
| wnt2 | 0.120 | alyref | -0.026 |
| si:dkey-247m21.3 | 0.119 | hmgb1b | -0.026 |
| triobpa | 0.118 | hnrnpabb | -0.026 |
| cracr2ab | 0.117 | h3f3a | -0.026 |
| tspan10 | 0.117 | apex1 | -0.026 |