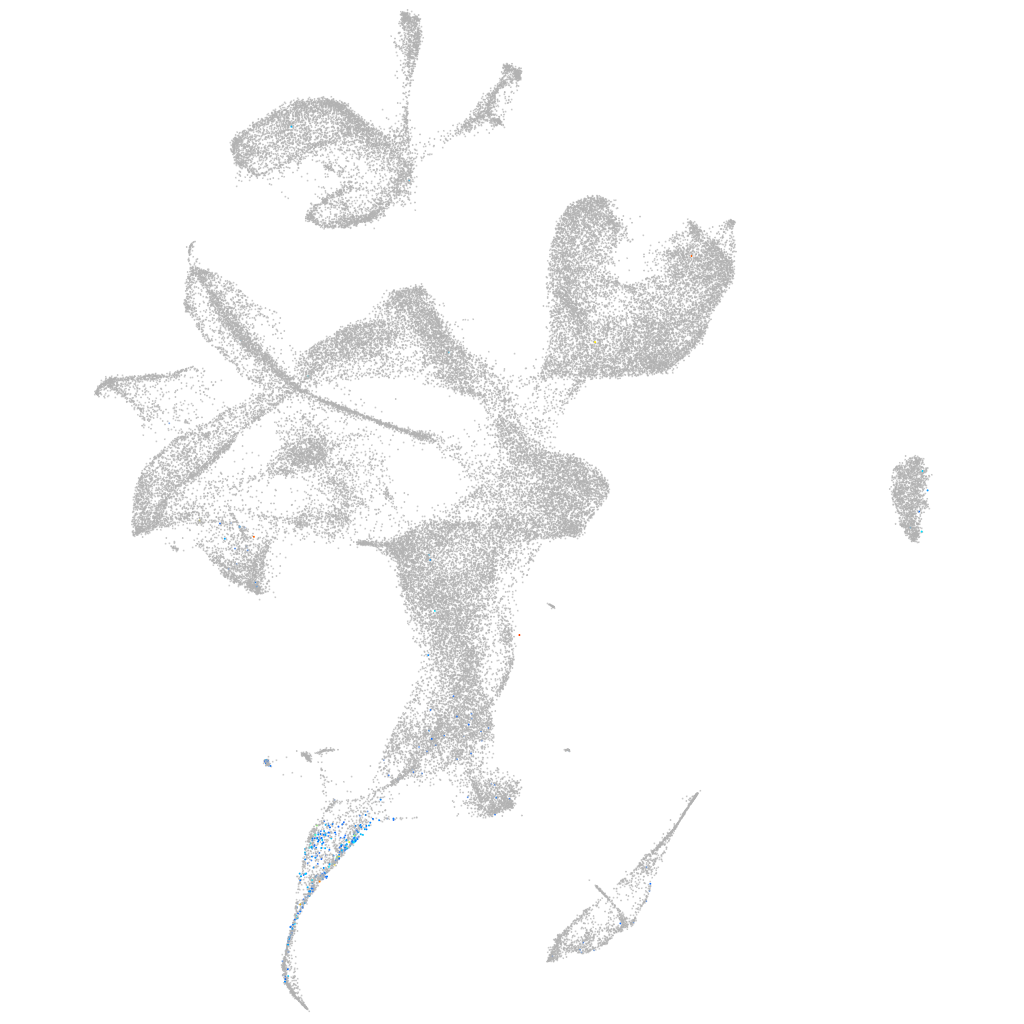
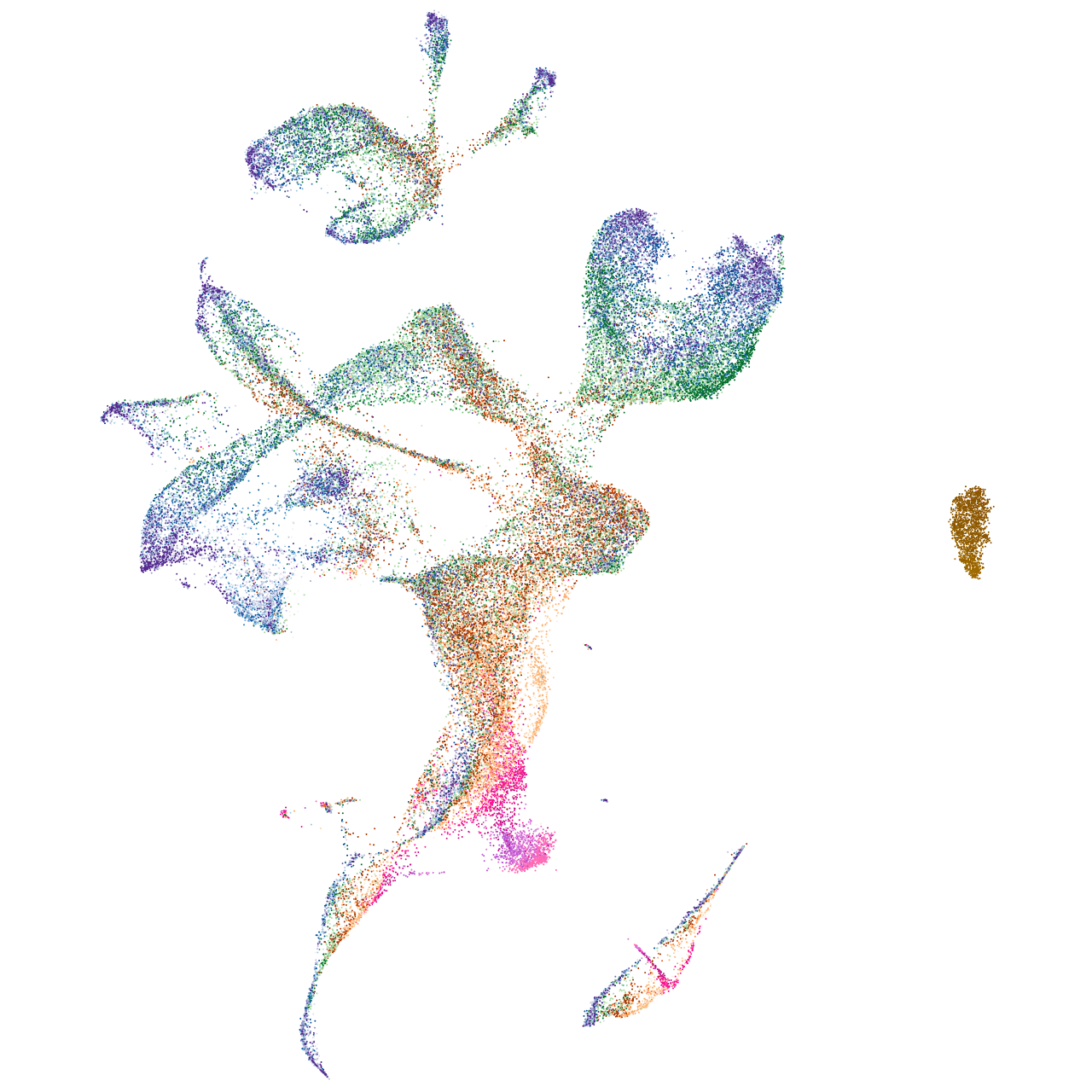sorting nexin 8a
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| slc45a2 | 0.285 | ptmab | -0.101 |
| gpr143 | 0.283 | ptmaa | -0.078 |
| rab32a | 0.261 | si:ch73-1a9.3 | -0.078 |
| dct | 0.258 | tuba1c | -0.072 |
| agtrap | 0.256 | gpm6aa | -0.071 |
| mitfa | 0.254 | hmgb1a | -0.062 |
| FP085398.1 | 0.254 | si:ch211-137a8.4 | -0.061 |
| qdpra | 0.250 | nova2 | -0.052 |
| rab38 | 0.250 | cadm3 | -0.051 |
| tspan36 | 0.248 | hmgn6 | -0.051 |
| zgc:110591 | 0.245 | tuba1a | -0.048 |
| tmem88b | 0.245 | epb41a | -0.047 |
| pmela | 0.242 | gpm6ab | -0.047 |
| tyrp1b | 0.241 | fabp7a | -0.046 |
| pnp4a | 0.237 | ckbb | -0.045 |
| pcbd1 | 0.235 | anp32e | -0.045 |
| sytl2b | 0.234 | hmgb3a | -0.045 |
| tyrp1a | 0.234 | hmgb1b | -0.045 |
| slc24a5 | 0.230 | rorb | -0.045 |
| tspan10 | 0.227 | foxg1b | -0.043 |
| pmelb | 0.222 | mdkb | -0.043 |
| tfec | 0.220 | snap25b | -0.043 |
| tyr | 0.218 | rtn1a | -0.041 |
| oca2 | 0.217 | stmn1b | -0.041 |
| gstp1 | 0.212 | gapdhs | -0.040 |
| bace2 | 0.209 | marcksl1b | -0.040 |
| pah | 0.206 | si:dkey-276j7.1 | -0.039 |
| alkal2a | 0.201 | si:ch1073-429i10.3.1 | -0.039 |
| cracr2ab | 0.197 | neurod4 | -0.039 |
| trpm1b | 0.193 | rnasekb | -0.039 |
| cx43 | 0.190 | crx | -0.038 |
| rnaseka | 0.190 | cspg5a | -0.038 |
| wnt2 | 0.188 | baz2ba | -0.038 |
| amdhd2 | 0.187 | CR383676.1 | -0.037 |
| triobpa | 0.184 | ndrg4 | -0.036 |