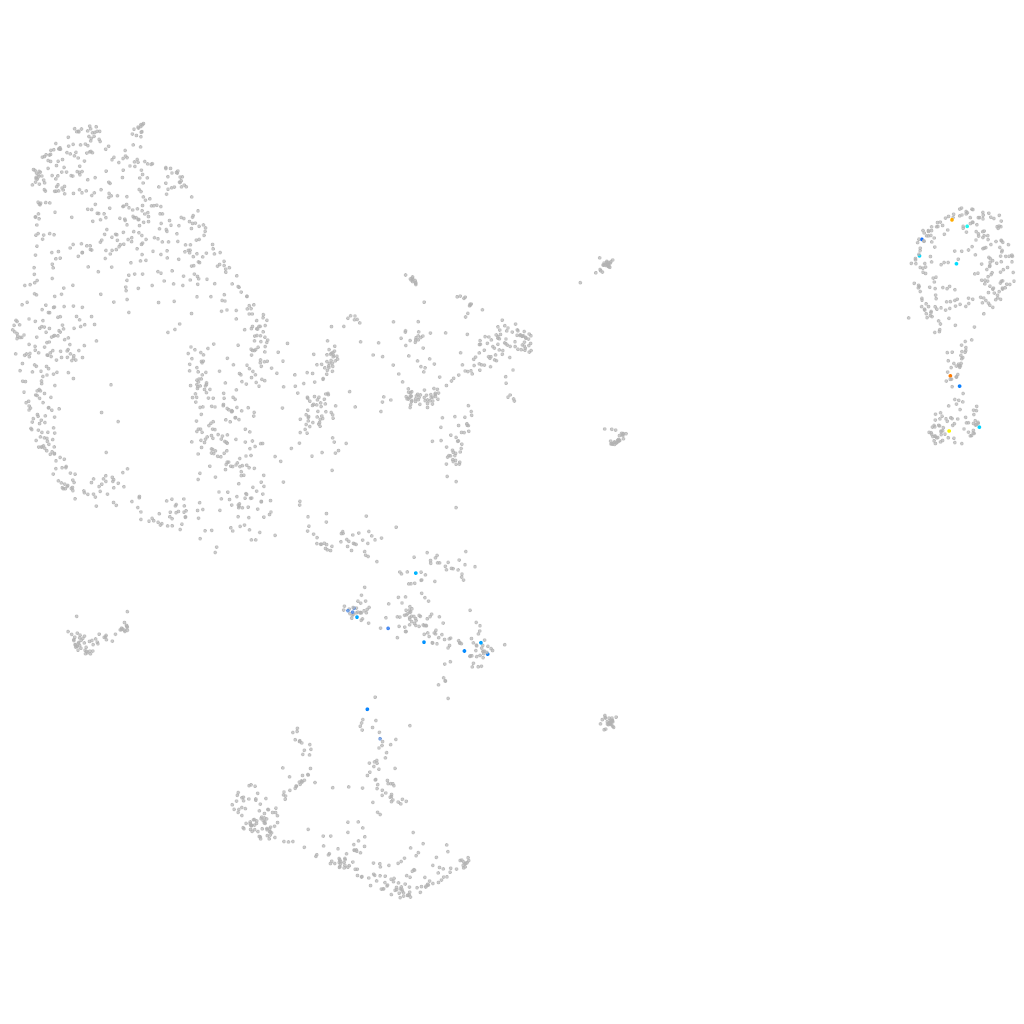
si:zfos-1505d6.3
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| zp2.3 | 0.420 | rpl37 | -0.112 |
| batf | 0.353 | atp5f1b | -0.111 |
| lrrn3a | 0.331 | si:ch211-139a5.9 | -0.107 |
| CU571069.1 | 0.276 | mt-atp6 | -0.106 |
| BX942837.1 | 0.267 | rps10 | -0.104 |
| si:dkeyp-82a1.4 | 0.267 | mt-cyb | -0.103 |
| adipoqa | 0.248 | atp5fa1 | -0.103 |
| znf1099 | 0.248 | vdac3 | -0.102 |
| guca1c | 0.239 | atp1b1a | -0.102 |
| AL928908.2 | 0.236 | eno3 | -0.102 |
| CT030005.2 | 0.236 | zgc:114188 | -0.100 |
| XLOC-012946 | 0.236 | ldhba | -0.097 |
| hsd3b1 | 0.231 | COX3 | -0.096 |
| zp3c | 0.231 | ndufa3 | -0.095 |
| LOC103911487 | 0.226 | mt-co2 | -0.094 |
| zp2.1 | 0.226 | rps17 | -0.092 |
| si:dkey-261m9.6 | 0.225 | prdx2 | -0.092 |
| mcoln3a | 0.224 | atp5pf | -0.091 |
| XLOC-040516 | 0.213 | suclg1 | -0.090 |
| CR788230.1 | 0.207 | cox6a1 | -0.089 |
| XLOC-005216 | 0.204 | mt-co1 | -0.089 |
| dre-let-7g-1 | 0.199 | atp5mc3b | -0.088 |
| serpinb1l4 | 0.198 | aldh6a1 | -0.088 |
| lygl2 | 0.197 | atp5mc1 | -0.087 |
| nr0b1 | 0.197 | atp5f1e | -0.087 |
| CU694380.1 | 0.196 | mt-nd4 | -0.086 |
| si:dkey-90l23.1 | 0.195 | mdh1aa | -0.084 |
| gas2l1 | 0.194 | mt-nd2 | -0.083 |
| trpc4apb | 0.191 | cox6b1 | -0.082 |
| cgas | 0.190 | atp5pd | -0.082 |
| TCIM | 0.189 | tpi1b | -0.082 |
| ifit11 | 0.185 | mt-nd1 | -0.082 |
| zwilch | 0.183 | GCA | -0.082 |
| bscl2l | 0.180 | elovl1b | -0.081 |
| abraxas1 | 0.179 | cotl1 | -0.081 |