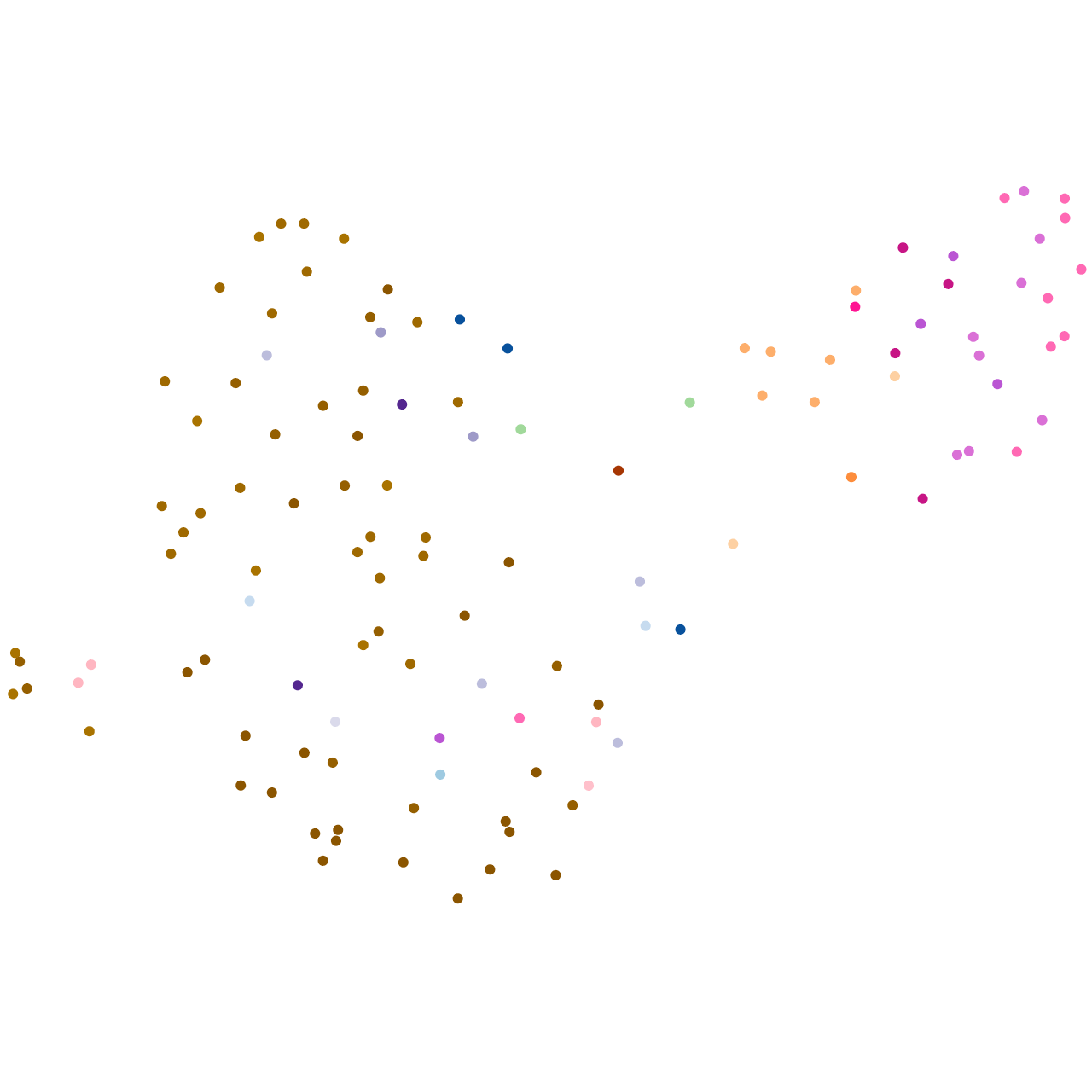
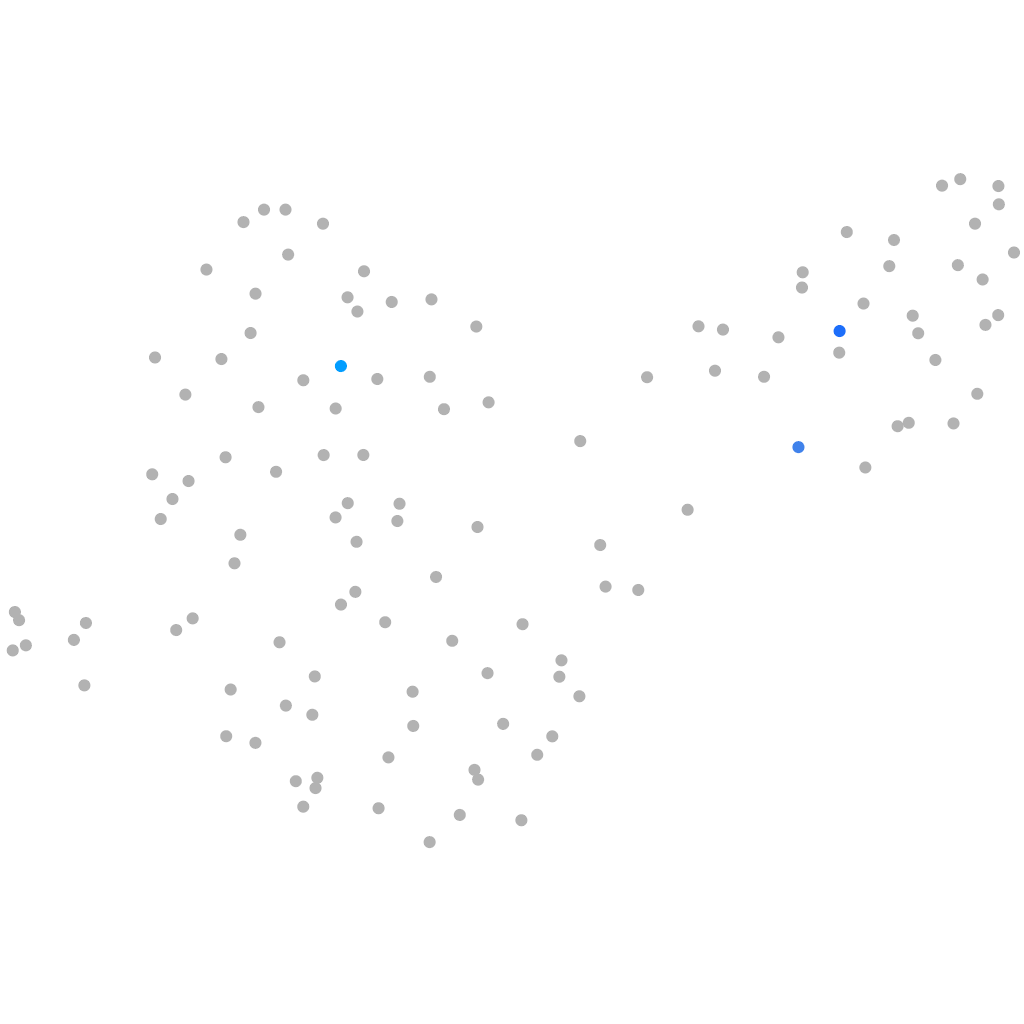
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| ugt1b5 | 0.879 | sf3b2 | -0.251 |
| efr3ba | 0.794 | hnrnpa1a | -0.202 |
| emilin2a | 0.794 | nme2b.1 | -0.202 |
| kif3cb | 0.794 | ipo7 | -0.194 |
| si:ch1073-513e17.1 | 0.794 | gar1 | -0.191 |
| si:dkey-1h24.2 | 0.794 | puf60b | -0.187 |
| pnp5b | 0.778 | tubb2b | -0.183 |
| cenpv | 0.722 | sec62 | -0.182 |
| sept4b | 0.683 | ddx18 | -0.180 |
| LOC108181079 | 0.681 | fnbp4 | -0.178 |
| CU695223.1 | 0.674 | capza1b | -0.177 |
| hoxa13b | 0.674 | srrt | -0.176 |
| LOC108181166 | 0.666 | myl12.1 | -0.175 |
| kita | 0.649 | snrpa1 | -0.173 |
| CT025771.1 | 0.646 | ddx42 | -0.170 |
| exoc3l1 | 0.639 | apex1 | -0.169 |
| fam171b | 0.616 | u2af2a | -0.168 |
| pdss2 | 0.606 | cdv3 | -0.167 |
| pdlim4 | 0.600 | rnf40 | -0.165 |
| filip1l | 0.597 | coil | -0.161 |
| arhgap36 | 0.596 | ssrp1a | -0.158 |
| si:ch73-89b15.3 | 0.591 | srrm2 | -0.156 |
| LOC100537032 | 0.588 | psmd7 | -0.155 |
| LOC100535210 | 0.583 | sap18 | -0.152 |
| disp1 | 0.574 | spns1 | -0.151 |
| trim62 | 0.569 | gapdh | -0.147 |
| bin1a | 0.563 | kctd5a | -0.147 |
| ano1 | 0.545 | ewsr1a | -0.146 |
| LOC101884382 | 0.539 | senp3b | -0.146 |
| LOC110438587 | 0.534 | hp1bp3 | -0.145 |
| slc4a1b | 0.529 | ncaph | -0.144 |
| camkva | 0.522 | si:ch211-212k18.4 | -0.143 |
| znf692 | 0.518 | si:ch211-173p18.3 | -0.143 |
| ccdc96 | 0.512 | psmb4 | -0.142 |
| stmn2a | 0.512 | slc25a5 | -0.140 |