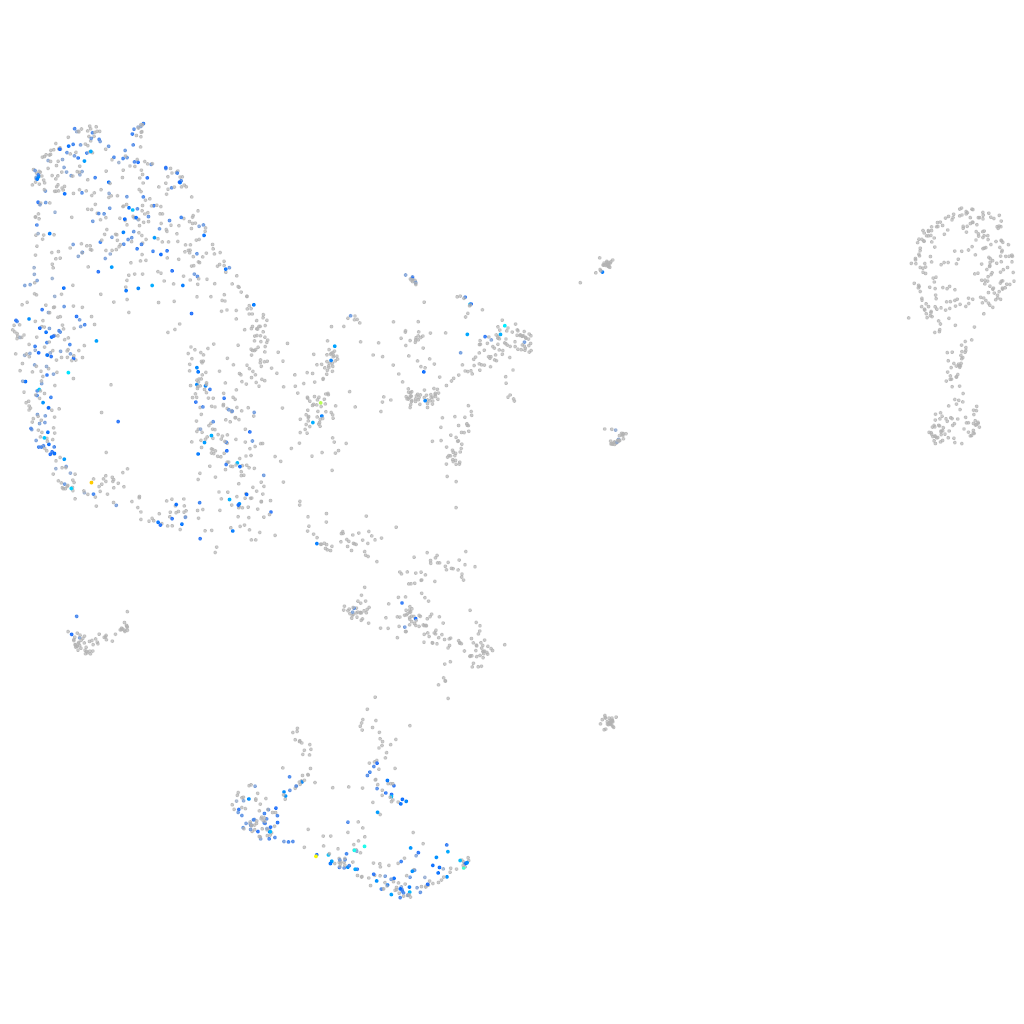
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| chp1 | 0.293 | hmga1a | -0.220 |
| cox7a1 | 0.292 | si:ch73-281n10.2 | -0.219 |
| cox6a2 | 0.284 | hnrnpabb | -0.206 |
| sod2 | 0.283 | hmgb2b | -0.205 |
| cdaa | 0.281 | h2afvb | -0.205 |
| ldhba | 0.270 | hspb1 | -0.202 |
| sstr5 | 0.269 | si:ch73-1a9.3 | -0.201 |
| cdh17 | 0.267 | h3f3d | -0.201 |
| glud1a | 0.267 | marcksb | -0.200 |
| atp1b1a | 0.266 | si:ch211-222l21.1 | -0.196 |
| slc25a5 | 0.265 | nucks1a | -0.194 |
| si:ch211-139a5.9 | 0.259 | hmgn2 | -0.194 |
| acadm | 0.257 | marcksl1b | -0.190 |
| mt-nd4 | 0.255 | ptmab | -0.187 |
| atp5mc3b | 0.255 | wu:fb97g03 | -0.187 |
| si:dkey-16p21.8 | 0.255 | msna | -0.182 |
| chchd10 | 0.255 | apoeb | -0.181 |
| bin2a | 0.252 | seta | -0.180 |
| vill | 0.249 | syncrip | -0.178 |
| sgk2a | 0.248 | hmgb2a | -0.178 |
| echs1 | 0.248 | setb | -0.177 |
| trim35-12 | 0.248 | hmgb1b | -0.174 |
| mt-atp6 | 0.242 | hmgn6 | -0.172 |
| mt-nd1 | 0.242 | hnrnph1l | -0.169 |
| atp5f1b | 0.241 | hnrnpaba | -0.168 |
| uqcrc1 | 0.240 | hnrnpa0b | -0.166 |
| atp5pd | 0.237 | akap12b | -0.165 |
| ponzr1 | 0.237 | ilf3b | -0.165 |
| mt-nd2 | 0.234 | acin1a | -0.165 |
| atp5f1c | 0.233 | apoc1 | -0.165 |
| atp5fa1 | 0.233 | tubb2b | -0.164 |
| mdh1aa | 0.232 | khdrbs1a | -0.163 |
| atp5pf | 0.232 | si:ch211-288g17.3 | -0.162 |
| gstk1 | 0.231 | zgc:110425 | -0.162 |
| slc7a8a | 0.231 | anp32a | -0.160 |