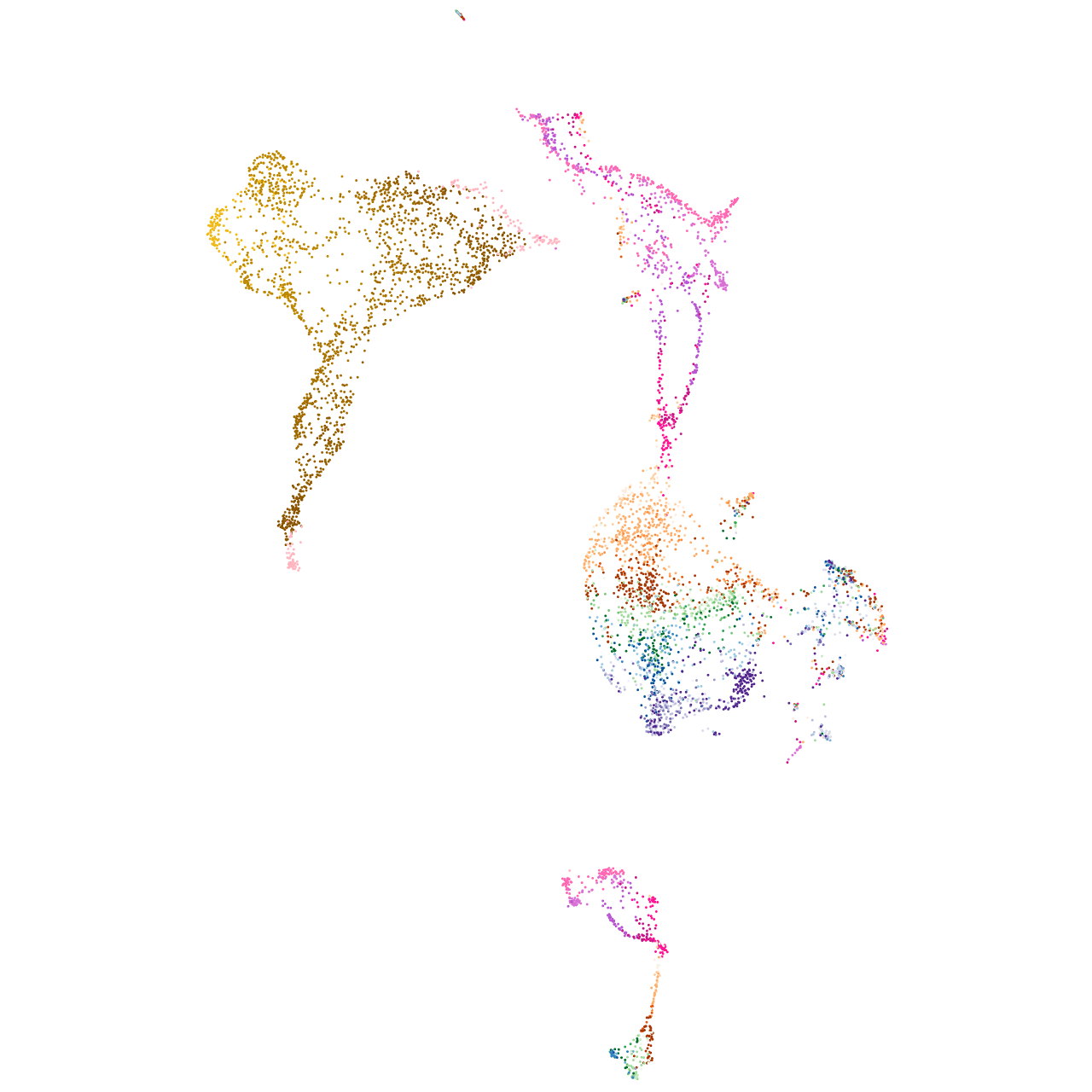paired box 5
ZFIN 
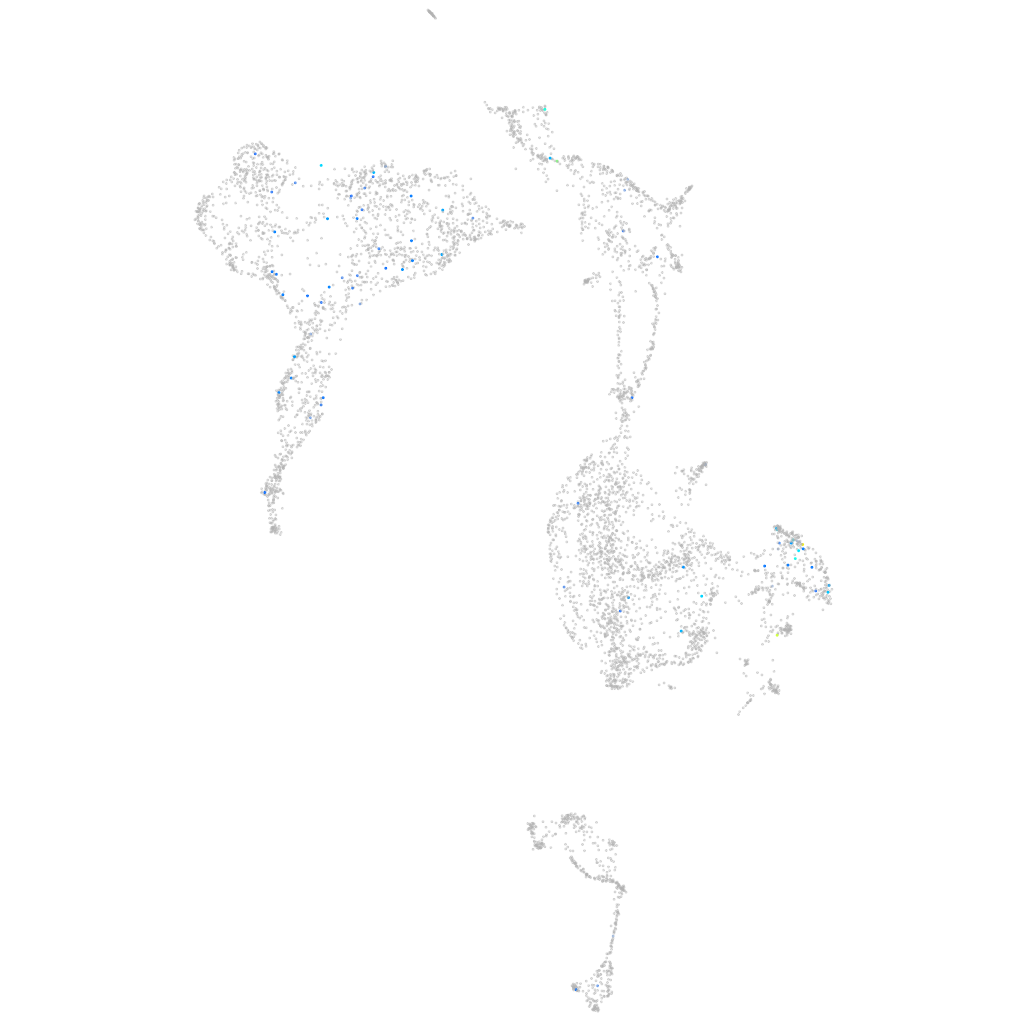
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| en2b | 0.282 | fkbp11 | -0.046 |
| LOC110438540 | 0.254 | krt18a.1 | -0.042 |
| pax2a | 0.251 | zgc:153675 | -0.041 |
| pax8 | 0.220 | anxa13 | -0.041 |
| aqp4 | 0.215 | rpl8 | -0.040 |
| tal1 | 0.214 | pdcd6 | -0.040 |
| en2a | 0.202 | cav2 | -0.040 |
| en1b | 0.192 | tpm4a | -0.040 |
| zgc:171470 | 0.185 | ssr2 | -0.040 |
| CABZ01084566.1 | 0.183 | fxyd1 | -0.040 |
| si:ch73-233f7.3 | 0.181 | haus7 | -0.039 |
| pax2b | 0.171 | zgc:162730 | -0.039 |
| otofa | 0.167 | krt8 | -0.039 |
| sox14 | 0.166 | atox1 | -0.038 |
| igfbp1b | 0.158 | col8a1a | -0.038 |
| XLOC-039965 | 0.153 | cybrd1 | -0.038 |
| pcdh1gb2 | 0.151 | copz2 | -0.038 |
| prelp | 0.150 | serp1 | -0.038 |
| XLOC-019859 | 0.149 | cyr61 | -0.038 |
| INSM2 | 0.148 | fam114a1 | -0.038 |
| pcdh2g5 | 0.146 | cav1 | -0.037 |
| CU013548.1 | 0.145 | smco4 | -0.037 |
| LOC101886244 | 0.144 | anxa11a | -0.037 |
| sox21a | 0.142 | zgc:113921 | -0.036 |
| si:ch211-285j22.3.1 | 0.139 | slc3a2a | -0.036 |
| sult4a1 | 0.139 | ptk2ab | -0.036 |
| pou3f2b | 0.136 | anxa2a | -0.036 |
| LOC103909470 | 0.134 | rps20 | -0.036 |
| XLOC-022007 | 0.134 | kdelr3 | -0.036 |
| CU633762.1 | 0.134 | rgcc | -0.035 |
| XLOC-043772 | 0.134 | slc30a7 | -0.035 |
| oprk1 | 0.134 | cd164 | -0.035 |
| XLOC-003053 | 0.132 | ckap4 | -0.035 |
| spock1 | 0.132 | mcfd2 | -0.035 |
| emp1 | 0.131 | kdelr2b | -0.035 |