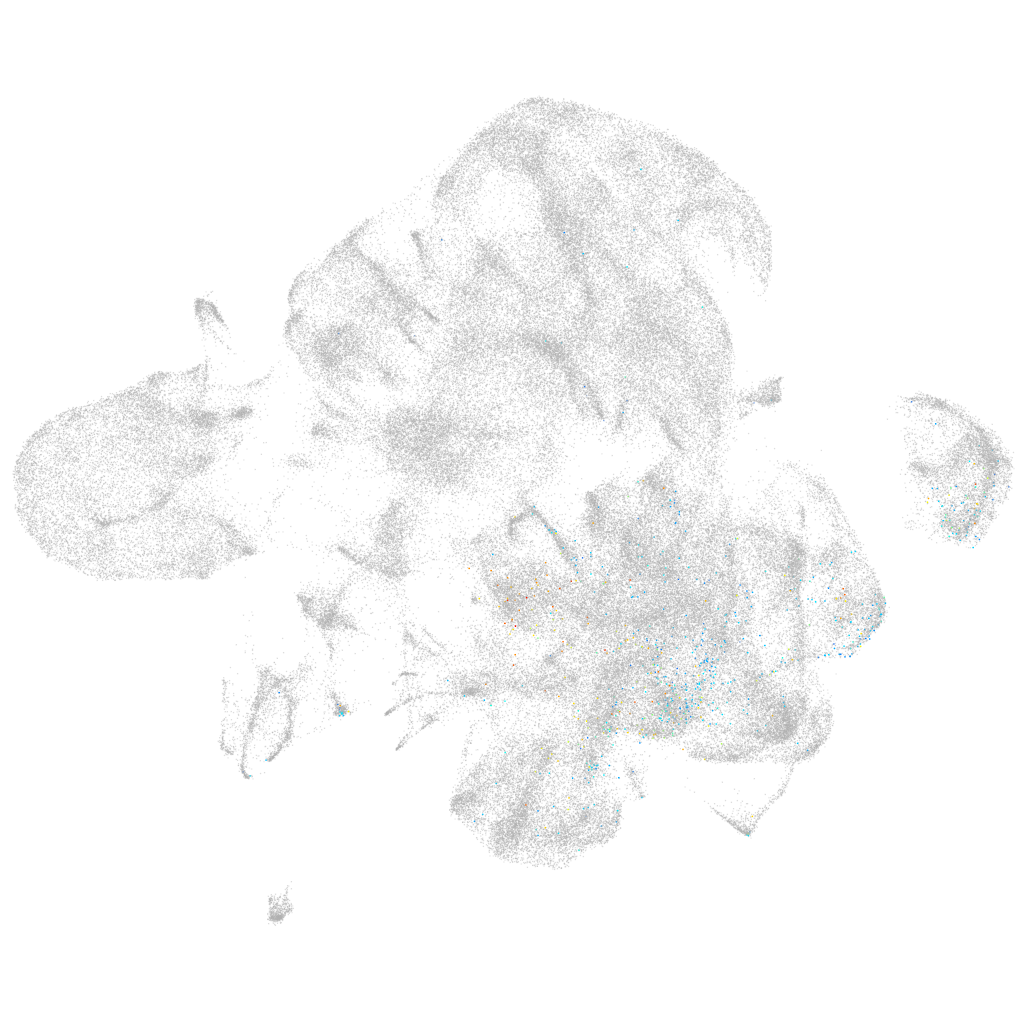
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm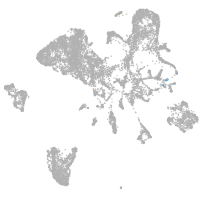 |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle 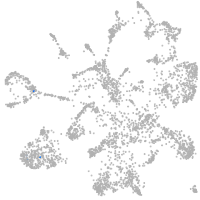 |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural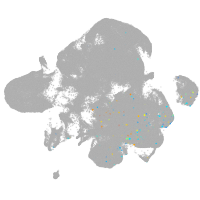 |
spinal cord / glia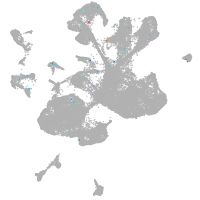 |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| CABZ01043955.1 | 0.117 | si:dkey-151g10.6 | -0.073 |
| scg2b | 0.108 | rplp2l | -0.065 |
| atp6v0cb | 0.088 | rps28 | -0.063 |
| ywhag2 | 0.082 | hmgb2b | -0.063 |
| sypb | 0.081 | rplp1 | -0.062 |
| sncb | 0.079 | hmga1a | -0.060 |
| vamp2 | 0.078 | rpl39 | -0.059 |
| atpv0e2 | 0.077 | rps20 | -0.058 |
| rnasekb | 0.076 | hmgb2a | -0.058 |
| rtn1b | 0.076 | rps12 | -0.058 |
| camk2n1a | 0.075 | rpl36a | -0.056 |
| gpm6ab | 0.073 | rpl26 | -0.056 |
| spock3 | 0.073 | eef1a1l1 | -0.056 |
| atp6ap2 | 0.073 | rpl23 | -0.056 |
| stmn2a | 0.072 | rpl11 | -0.056 |
| gapdhs | 0.072 | rps19 | -0.055 |
| syt1a | 0.071 | rps7 | -0.055 |
| sv2a | 0.071 | rps29 | -0.055 |
| lingo2b | 0.070 | rpl12 | -0.054 |
| zgc:65894 | 0.070 | rpsa | -0.054 |
| gap43 | 0.070 | rps9 | -0.053 |
| gng3 | 0.069 | rps14 | -0.053 |
| sprn | 0.069 | ran | -0.053 |
| fxyd6 | 0.068 | rpl35a | -0.052 |
| syngr3a | 0.068 | rps2 | -0.052 |
| rab6bb | 0.068 | rpl10a | -0.052 |
| sypa | 0.068 | rpl38 | -0.051 |
| map1aa | 0.067 | rpl7a | -0.051 |
| si:dkeyp-75h12.5 | 0.067 | rps27a | -0.051 |
| serpini1 | 0.067 | rpl27 | -0.051 |
| si:dkeyp-72g9.4 | 0.067 | rpl21 | -0.050 |
| ndfip1 | 0.066 | rpl10 | -0.050 |
| snap25a | 0.066 | rps24 | -0.050 |
| dnajc5aa | 0.066 | rpl30 | -0.049 |
| gas7a | 0.066 | rpl18 | -0.049 |




