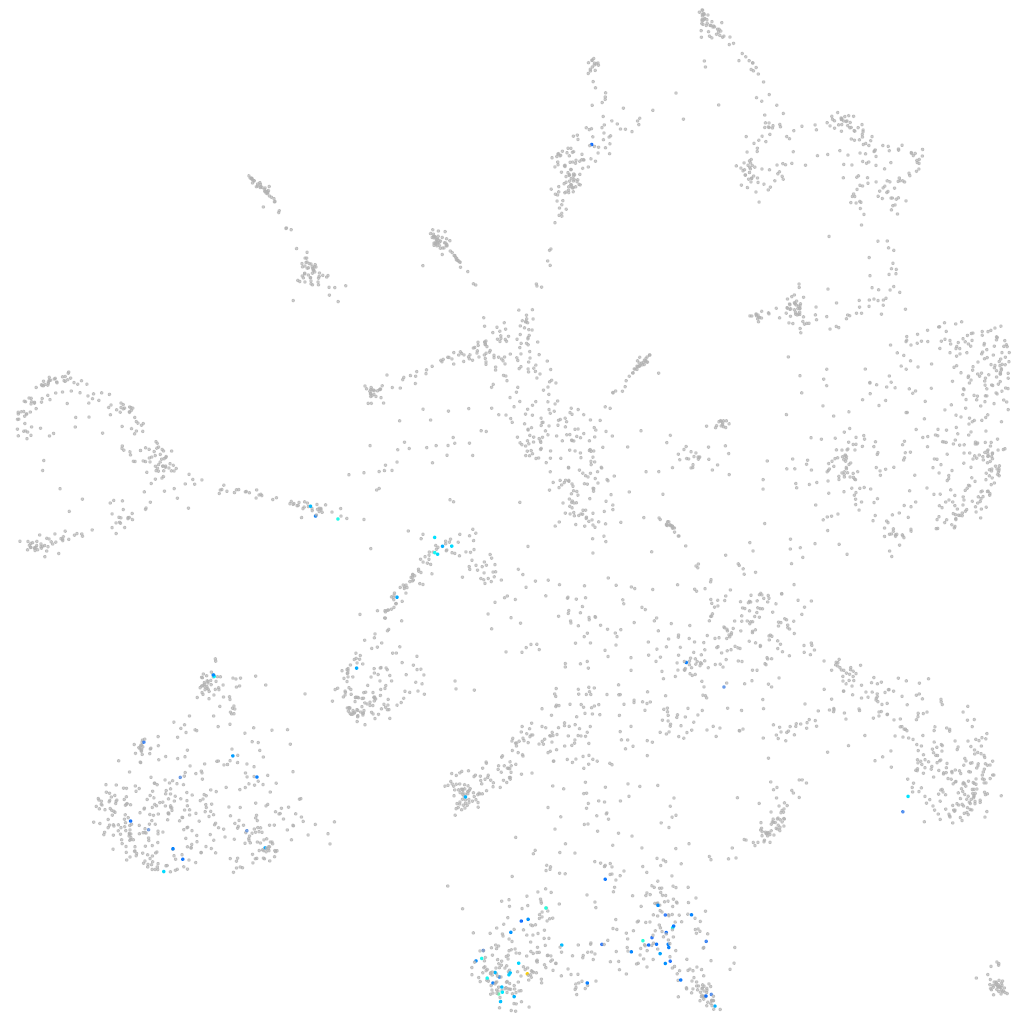
"guanine nucleotide binding protein (G protein), alpha 14a"
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| desmb | 0.243 | fthl27 | -0.072 |
| csrp1b | 0.235 | jam2b | -0.068 |
| cald1b | 0.225 | podxl | -0.064 |
| cnn1b | 0.220 | cd248a | -0.062 |
| tpm2 | 0.219 | krt94 | -0.062 |
| vcla | 0.217 | si:ch211-39i22.1 | -0.060 |
| fhl1a | 0.210 | tmem88b | -0.059 |
| fbxl22 | 0.209 | bambia | -0.059 |
| pdlim3a | 0.209 | cx43.4 | -0.058 |
| si:ch211-137i24.10 | 0.200 | nid2a | -0.058 |
| zgc:110699 | 0.194 | si:dkey-261h17.1 | -0.055 |
| mylkb | 0.188 | marcksl1b | -0.054 |
| myh11a | 0.186 | cpn1 | -0.054 |
| coro1a | 0.184 | lsp1 | -0.054 |
| grem1b | 0.178 | rbpms2b | -0.053 |
| rgs2 | 0.177 | ednraa | -0.052 |
| acta2 | 0.176 | tbx2b | -0.051 |
| smtna | 0.176 | serpine2 | -0.051 |
| XLOC-009861 | 0.175 | aqp1a.1 | -0.051 |
| tagln | 0.175 | cd151 | -0.051 |
| itpka | 0.172 | qkia | -0.050 |
| myl9a | 0.169 | sult6b1 | -0.049 |
| cabp1a | 0.168 | nid1a | -0.049 |
| LO018315.7 | 0.167 | bcam | -0.049 |
| ckbb | 0.166 | cxcl18b | -0.048 |
| nog2 | 0.165 | appa | -0.048 |
| akap7 | 0.164 | tubb2b | -0.047 |
| sorbs3 | 0.163 | hmga1a | -0.047 |
| XLOC-020459 | 0.162 | si:ch211-156j16.1 | -0.047 |
| tpm1 | 0.162 | tjp2b | -0.047 |
| myl6 | 0.162 | igf2b | -0.046 |
| CU468920.1 | 0.160 | add3a | -0.046 |
| acta1b | 0.159 | rpl38 | -0.046 |
| fhl5 | 0.159 | pdgfrb | -0.045 |
| LO018380.1 | 0.158 | cygb1 | -0.045 |