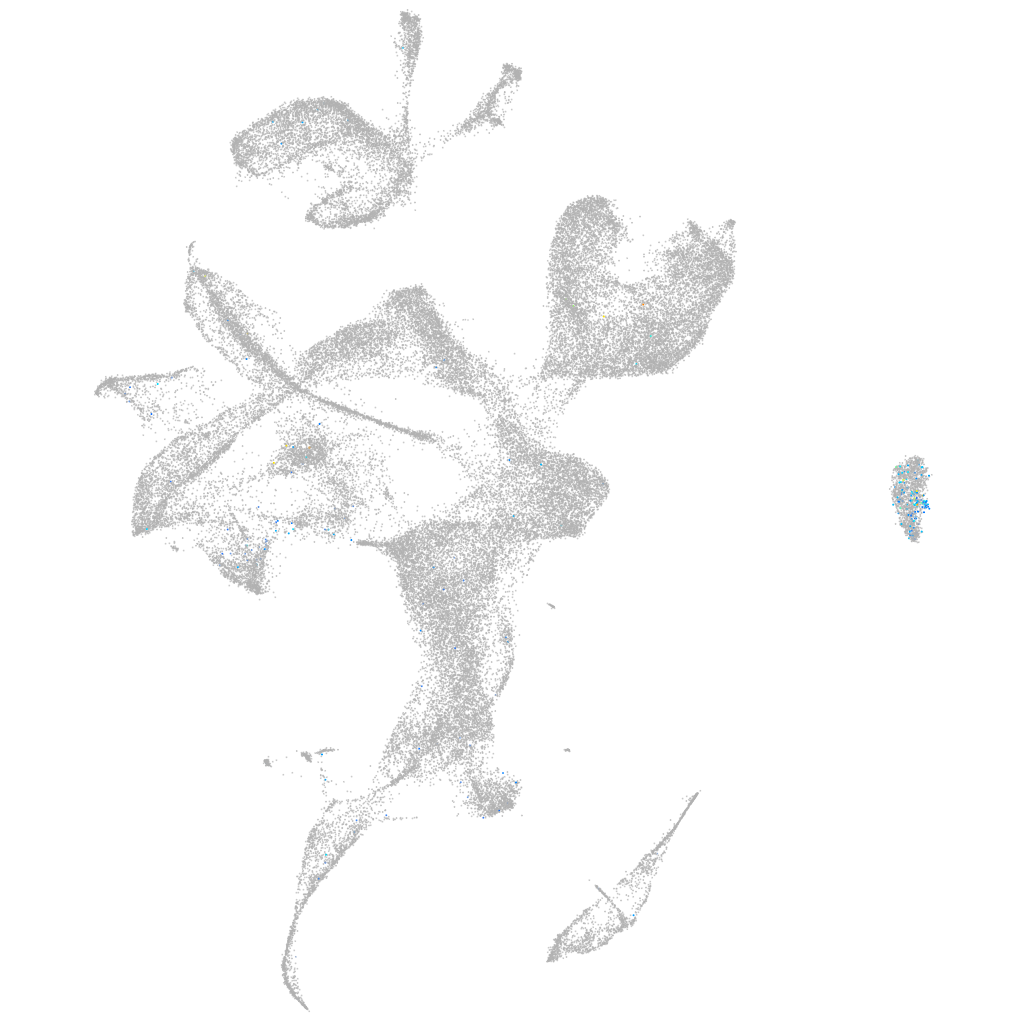
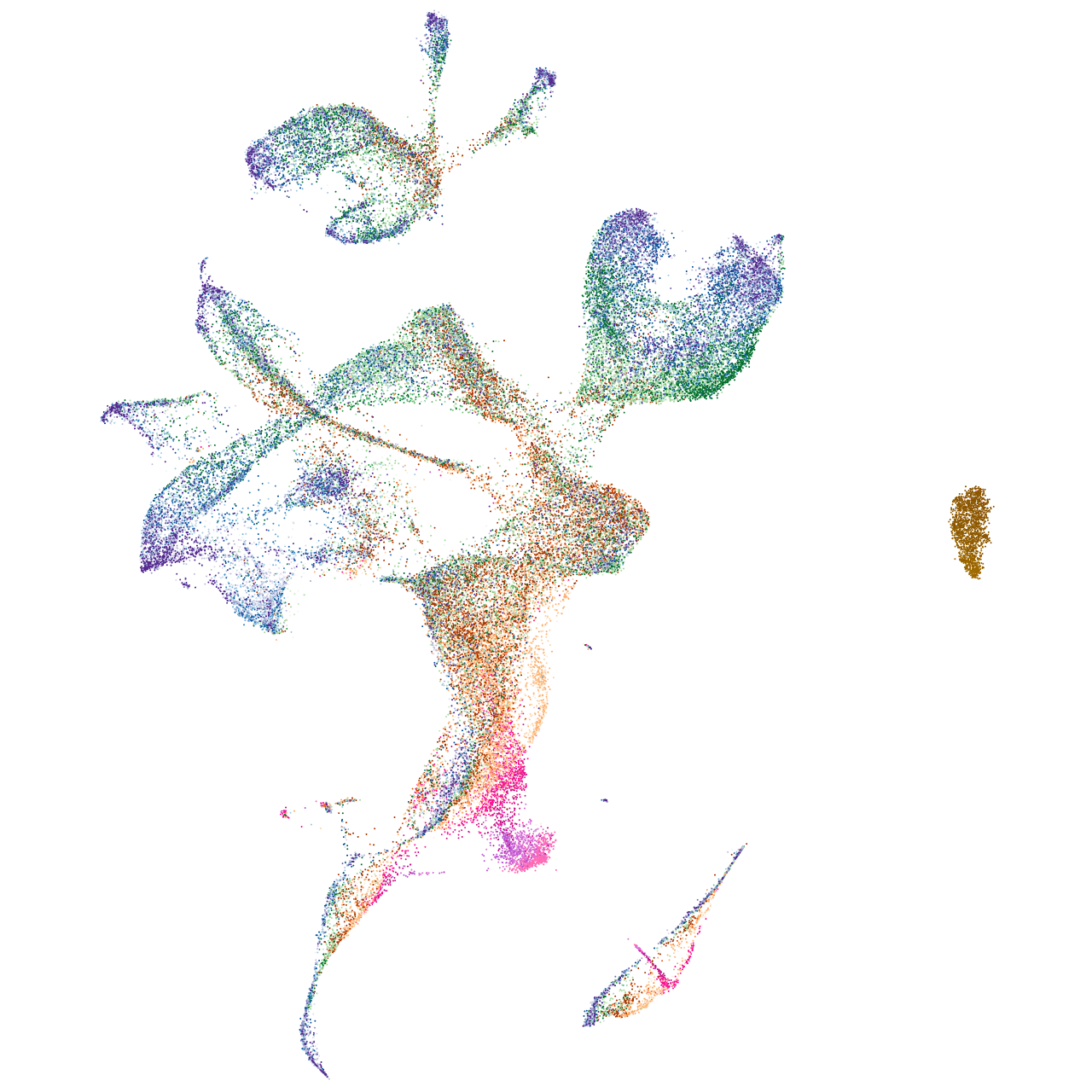forkhead box B1b
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| lft1 | 0.277 | rpl37 | -0.081 |
| nkx2.9 | 0.246 | rps10 | -0.080 |
| lft2 | 0.232 | zgc:114188 | -0.070 |
| foxd5 | 0.215 | h3f3a | -0.064 |
| sox19a | 0.181 | rps17 | -0.058 |
| ndr2 | 0.179 | ppiab | -0.053 |
| nkx2.4b | 0.174 | hnrnpa0l | -0.053 |
| gsc | 0.171 | ptmaa | -0.053 |
| nkx2.2b | 0.171 | si:ch1073-429i10.3.1 | -0.053 |
| dbx1a | 0.162 | CR383676.1 | -0.051 |
| tdgf1 | 0.161 | zgc:158463 | -0.049 |
| hspb1 | 0.159 | mt-nd1 | -0.043 |
| dact2 | 0.155 | COX3 | -0.043 |
| nr6a1a | 0.150 | mt-atp6 | -0.042 |
| chrd | 0.150 | mt-co2 | -0.041 |
| stm | 0.148 | cadm3 | -0.041 |
| foxb1a | 0.144 | ckbb | -0.040 |
| shisa2a | 0.144 | cct2 | -0.040 |
| ISCU | 0.138 | zgc:153409 | -0.040 |
| si:ch211-152c2.3 | 0.137 | atp5mc1 | -0.039 |
| admp | 0.135 | fabp7a | -0.038 |
| cyp26a1 | 0.126 | h3f3c | -0.035 |
| NAV1 | 0.123 | atp5f1b | -0.035 |
| LOC108179178 | 0.114 | gapdhs | -0.035 |
| COX7A2 | 0.113 | fabp3 | -0.034 |
| NC-002333.4 | 0.113 | tuba1c | -0.034 |
| pkdccb | 0.111 | mdh2 | -0.034 |
| hesx1 | 0.109 | marcksl1a | -0.034 |
| foxa | 0.107 | atp5mc3b | -0.033 |
| dnmt3bb.2 | 0.106 | pvalb1 | -0.033 |
| spic | 0.104 | atp6v1g1 | -0.033 |
| alcamb | 0.104 | atp5l | -0.033 |
| CU929403.2 | 0.104 | gpm6aa | -0.033 |
| nkx2.1 | 0.103 | hbae3 | -0.032 |
| si:ch211-176g6.2 | 0.103 | gpm6ab | -0.031 |