centrosomal protein 135
ZFIN 
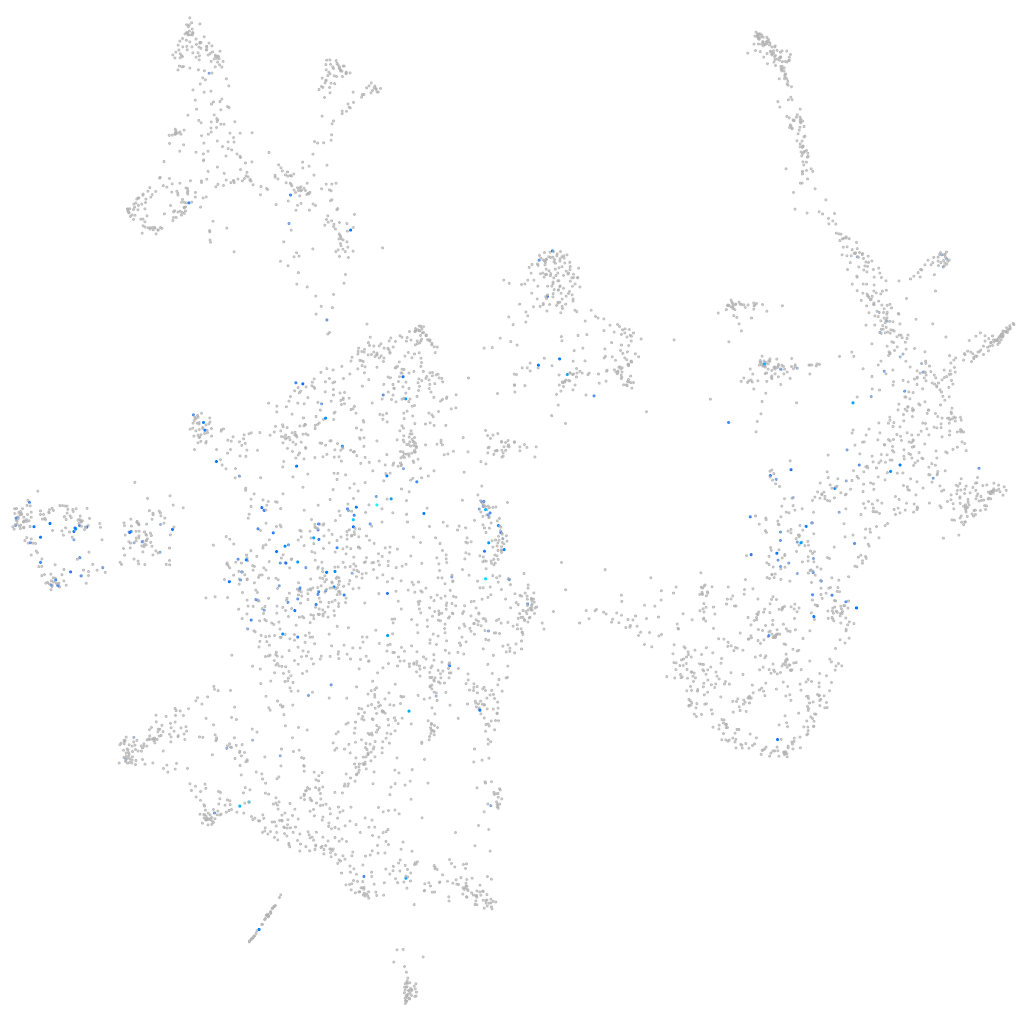
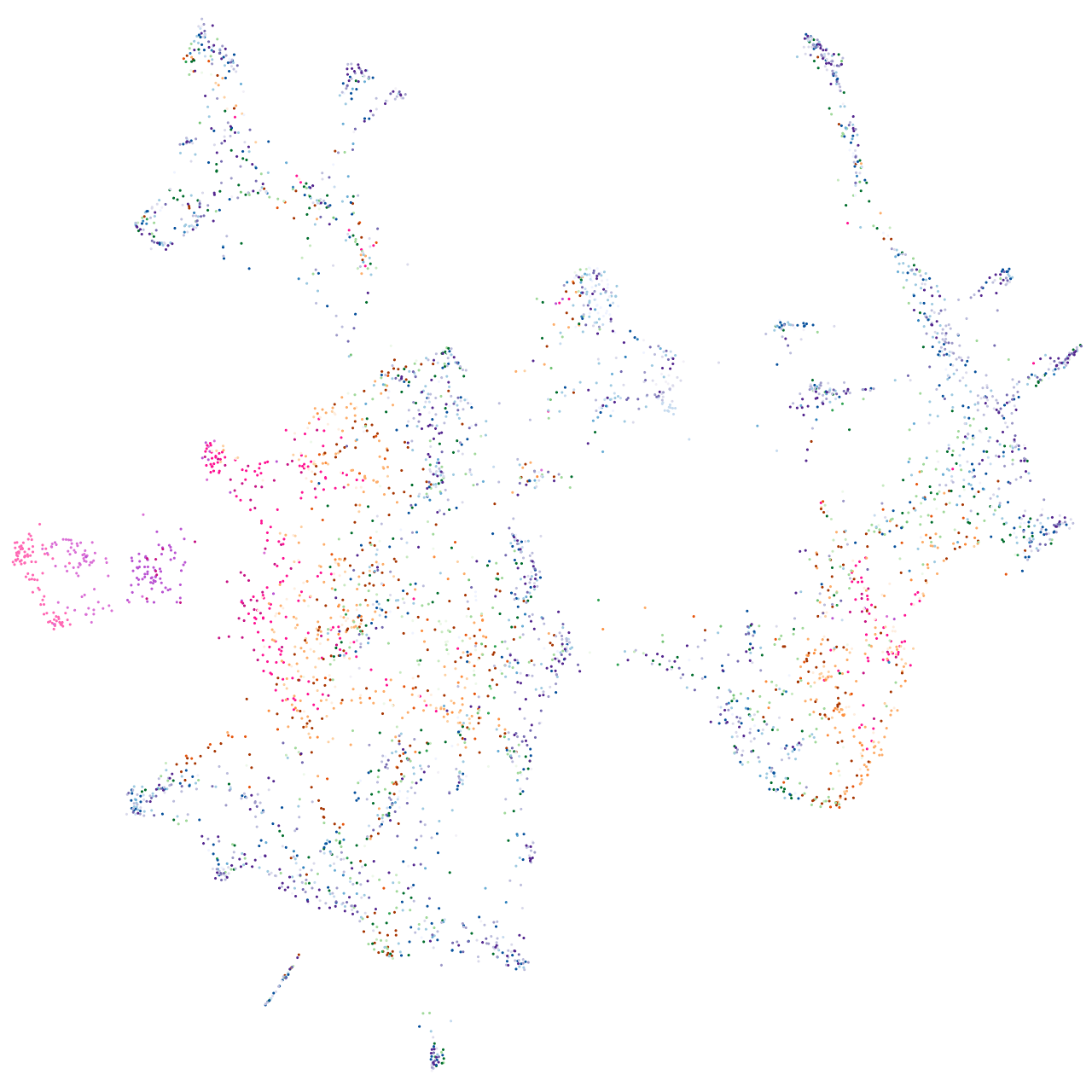
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| CABZ01005379.1 | 0.219 | pvalb8 | -0.078 |
| cks1b | 0.213 | calml4a | -0.074 |
| mibp | 0.212 | ap3s1.1 | -0.071 |
| mki67 | 0.207 | gapdhs | -0.070 |
| dut | 0.206 | osbpl1a | -0.070 |
| rrm2 | 0.202 | gpx2 | -0.070 |
| pcna | 0.201 | otofb | -0.069 |
| gins3 | 0.201 | pvalb1 | -0.067 |
| chaf1a | 0.194 | cd164l2 | -0.067 |
| ncapg | 0.192 | emb | -0.066 |
| dek | 0.191 | selenow2b | -0.065 |
| dhfr | 0.190 | atp1a3b | -0.064 |
| si:dkey-6i22.5 | 0.189 | pcsk5a | -0.064 |
| zgc:110540 | 0.189 | apba1a | -0.063 |
| ccna2 | 0.187 | si:dkey-17m8.1 | -0.062 |
| stmn1a | 0.187 | dnajc5b | -0.061 |
| tyms | 0.184 | syt14b | -0.061 |
| zgc:110216 | 0.184 | zgc:162989 | -0.061 |
| dctd | 0.182 | clic5b | -0.061 |
| prim2 | 0.181 | rnasekb | -0.061 |
| atad5a | 0.181 | si:dkey-16p21.8 | -0.060 |
| cdca5 | 0.178 | wnt2 | -0.060 |
| rrm1 | 0.178 | nptna | -0.060 |
| si:dkey-261m9.17 | 0.178 | CR925719.1 | -0.060 |
| mis12 | 0.177 | pkig | -0.060 |
| rpa2 | 0.174 | gb:eh456644 | -0.060 |
| zgc:165555.3 | 0.174 | eno1a | -0.060 |
| seta | 0.174 | calm1b | -0.059 |
| tk1 | 0.173 | atp2b1a | -0.059 |
| anp32b | 0.172 | si:ch73-15n15.3 | -0.059 |
| trip13 | 0.171 | lrrc73 | -0.059 |
| zgc:110425 | 0.170 | ip6k2a | -0.059 |
| tubb2b | 0.170 | ptbp3 | -0.059 |
| hmgb2a | 0.169 | prr15la | -0.058 |
| h2afx | 0.169 | pou4f3 | -0.058 |