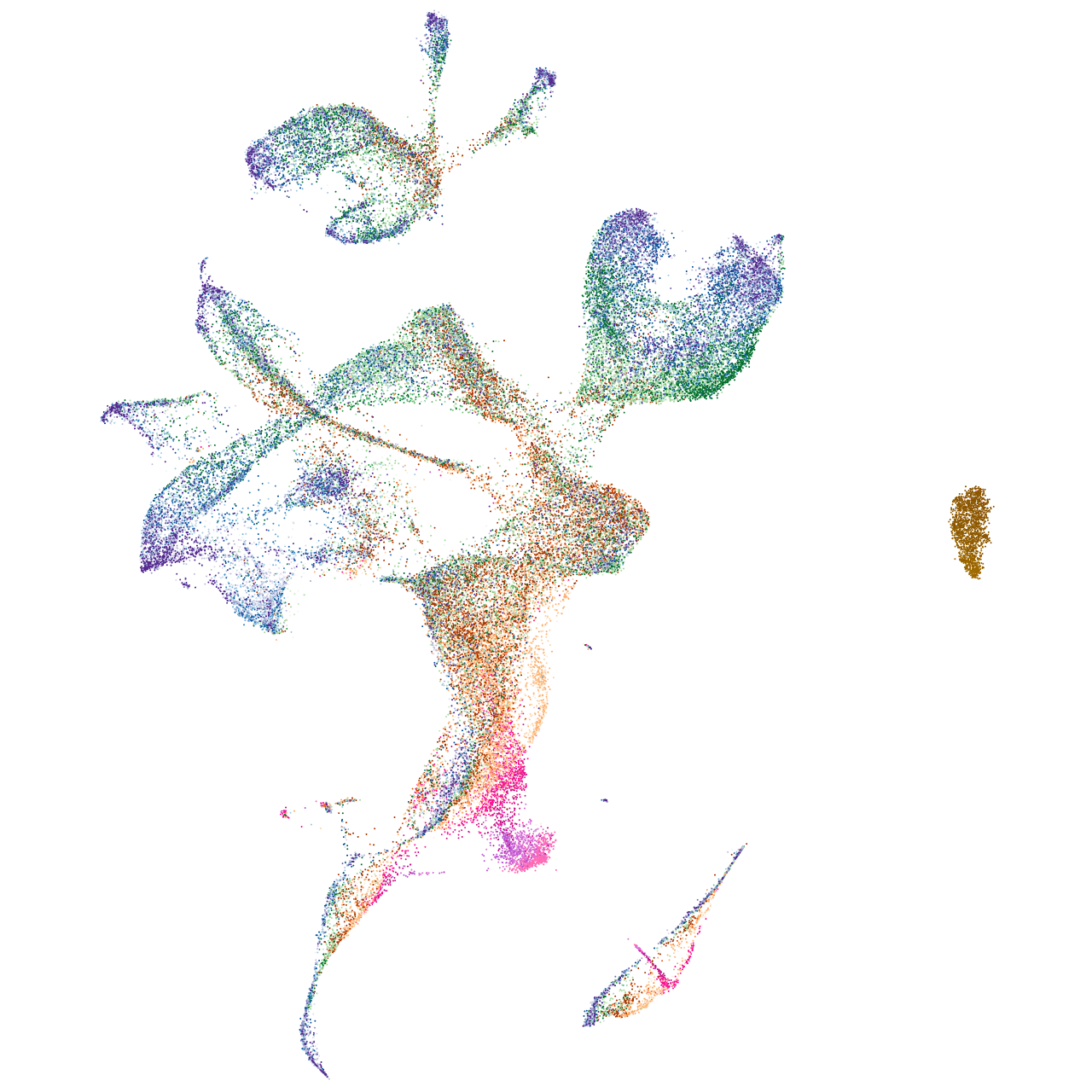C1q and TNF related 1
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| myh11a | 0.287 | ptmab | -0.047 |
| lmod1b | 0.285 | si:ch73-1a9.3 | -0.044 |
| rasl12 | 0.271 | marcksl1b | -0.035 |
| trh | 0.264 | khdrbs1a | -0.034 |
| cldn19 | 0.260 | si:ch211-137a8.4 | -0.034 |
| nmur1b | 0.259 | tuba1c | -0.032 |
| acta2 | 0.255 | hnrnpaba | -0.031 |
| thbs3b | 0.252 | marcksb | -0.031 |
| myocd | 0.249 | h2afva | -0.030 |
| wnt2 | 0.243 | cirbpb | -0.030 |
| mylka | 0.239 | hmga1a | -0.028 |
| ldlrap1a | 0.237 | fabp3 | -0.028 |
| FO744833.2 | 0.237 | snrpd1 | -0.027 |
| cldn7a | 0.229 | gpm6aa | -0.027 |
| sytl2b | 0.225 | tubb2b | -0.027 |
| pdlim3a | 0.222 | ckbb | -0.026 |
| tagln | 0.216 | cadm3 | -0.026 |
| artnb | 0.213 | hmgb1a | -0.026 |
| fhl2a | 0.206 | hmgn2 | -0.026 |
| fmn1 | 0.201 | fabp7a | -0.025 |
| hspg2 | 0.193 | snrpf | -0.025 |
| myl9a | 0.193 | tuba1a | -0.025 |
| BX088707.3 | 0.189 | gpm6ab | -0.024 |
| sorbs3 | 0.188 | hnrnpabb | -0.024 |
| fhl1a | 0.188 | hnrnpa0b | -0.024 |
| alkal2b | 0.186 | smarce1 | -0.023 |
| cnn1b | 0.182 | hmgb1b | -0.023 |
| malrd1 | 0.174 | nova2 | -0.023 |
| mlphb | 0.173 | ilf2 | -0.023 |
| myo1ca | 0.170 | epb41a | -0.023 |
| slc7a7 | 0.169 | snrpg | -0.023 |
| si:dkey-247m21.3 | 0.163 | ptges3b | -0.022 |
| BX323087.1 | 0.159 | hmgn6 | -0.022 |
| tyrp1b | 0.156 | anp32e | -0.022 |
| col9a1b | 0.154 | ddx39ab | -0.022 |