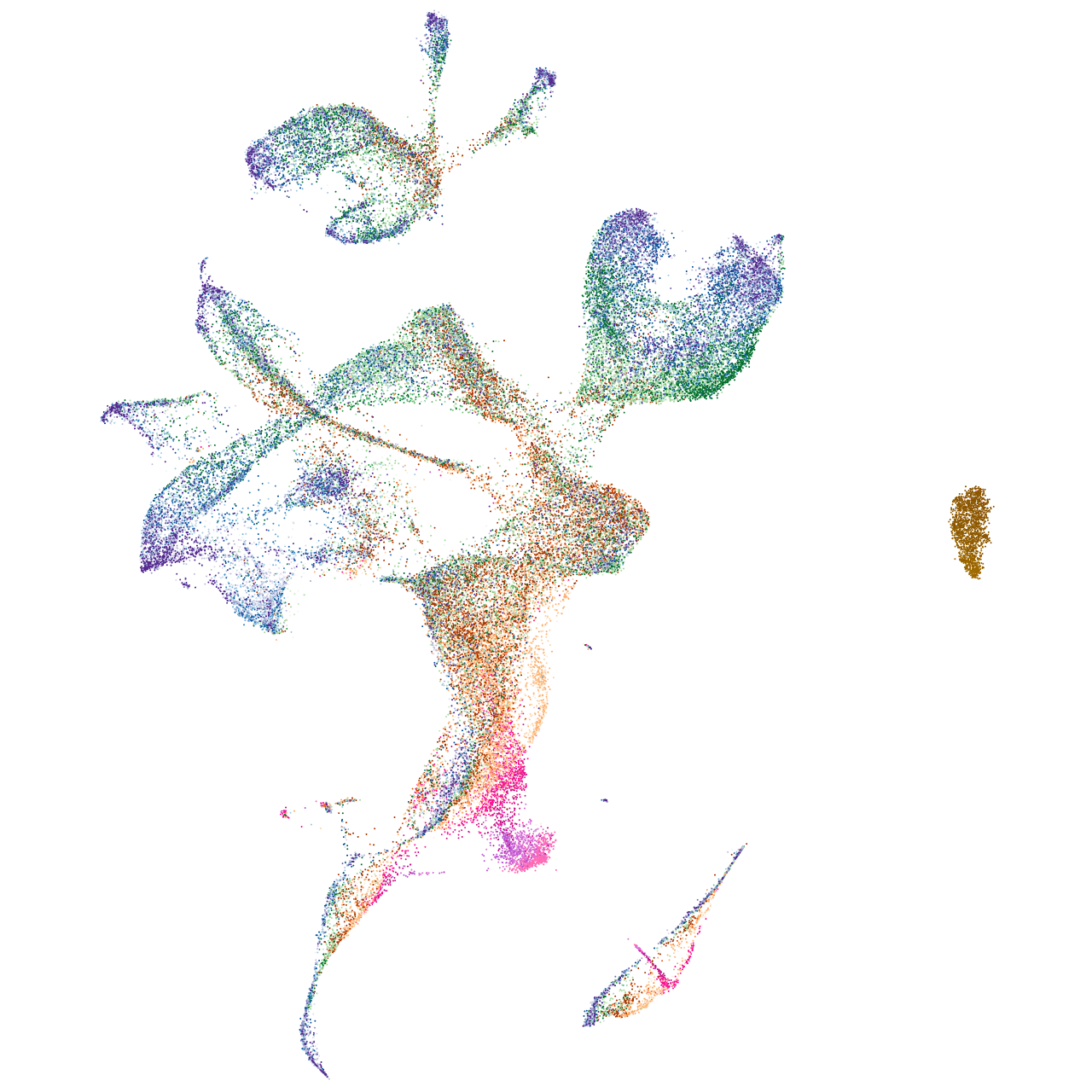Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| guca1b | 0.123 | si:ch211-222l21.1 | -0.043 |
| saga | 0.119 | cirbpb | -0.039 |
| sagb | 0.117 | hmgn2 | -0.036 |
| si:ch211-113d22.2 | 0.116 | marcksb | -0.034 |
| rom1b | 0.115 | hmgb1a | -0.033 |
| cnga1b | 0.114 | hnrnpaba | -0.032 |
| gnat1 | 0.114 | hmgb1b | -0.032 |
| cnga1a | 0.112 | cfl1 | -0.032 |
| gngt1 | 0.108 | hsp90ab1 | -0.032 |
| pde6b | 0.105 | hmga1a | -0.031 |
| pdca | 0.101 | marcksl1b | -0.031 |
| XLOC-002207 | 0.101 | hmgb2b | -0.031 |
| grk1a | 0.100 | khdrbs1a | -0.031 |
| aipl1 | 0.099 | h2afvb | -0.031 |
| CABZ01084566.1 | 0.099 | rpl9 | -0.029 |
| rbp2a | 0.098 | hnrnpa0b | -0.029 |
| BX004774.2 | 0.098 | nova2 | -0.028 |
| rho | 0.096 | hes6 | -0.027 |
| rhol | 0.096 | smarce1 | -0.027 |
| LOC101884626 | 0.095 | ran | -0.027 |
| guca1a | 0.093 | tubb2b | -0.027 |
| cplx4c | 0.092 | h3f3a | -0.026 |
| rom1a | 0.090 | sox11b | -0.026 |
| pde6ga | 0.086 | tuba1a | -0.025 |
| cabp4 | 0.084 | si:dkey-42i9.4 | -0.025 |
| zgc:77752 | 0.084 | marcksl1a | -0.025 |
| faimb | 0.083 | chd4a | -0.025 |
| unc119.2 | 0.081 | hnrnpa0l | -0.025 |
| zgc:112334 | 0.080 | ptmab | -0.025 |
| LOC110438506 | 0.079 | syncrip | -0.024 |
| zgc:162144 | 0.079 | rps16 | -0.024 |
| kcnv2a | 0.075 | actb1 | -0.024 |
| LOC100535225 | 0.073 | seta | -0.024 |
| LOC108180196 | 0.073 | hnrnpa1a | -0.024 |
| BX004873.1 | 0.073 | marcksa | -0.024 |