"bone morphogenetic protein receptor, type IBa"
ZFIN 
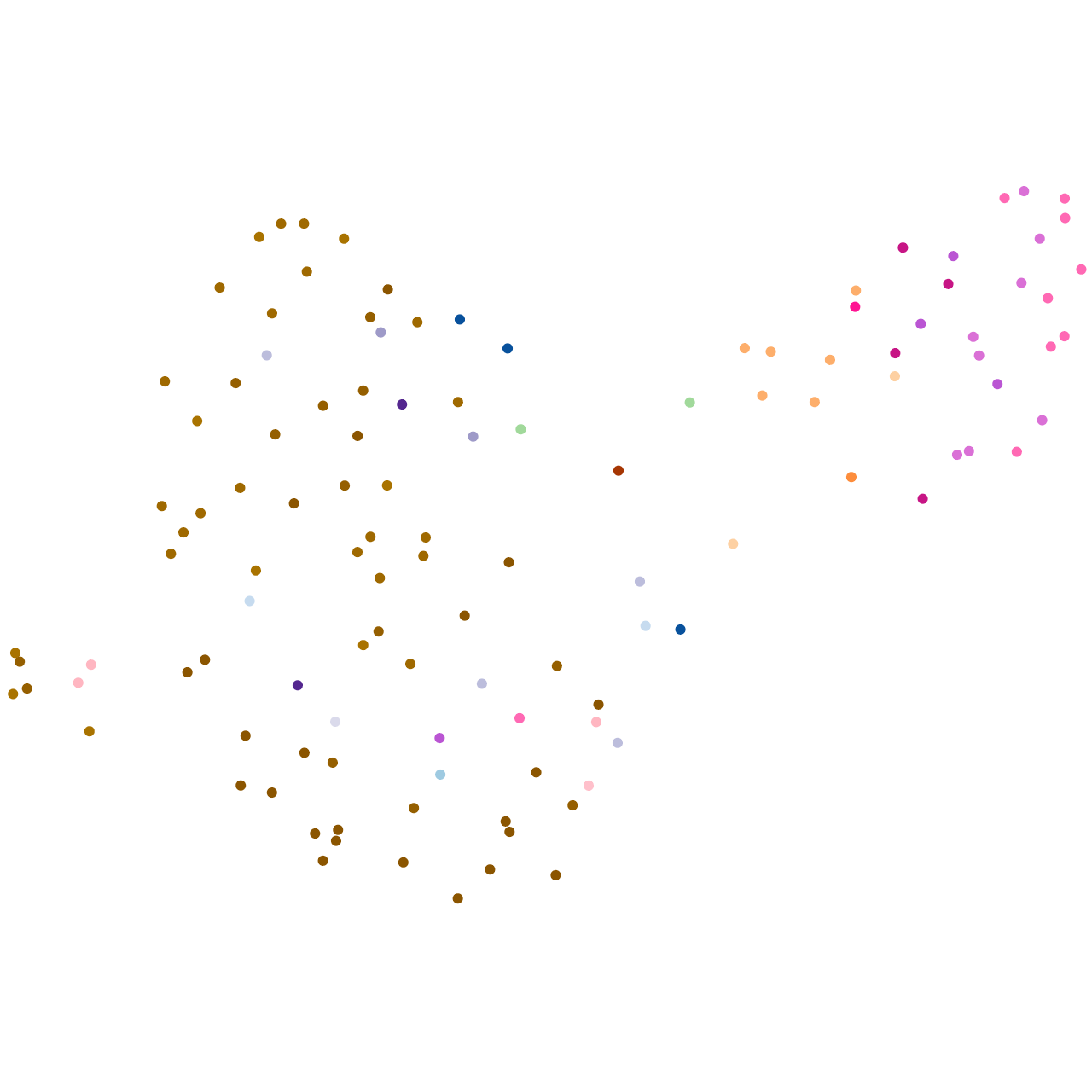
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| TCIM (1 of many) | 0.671 | hnrnpaba | -0.372 |
| FP102073.1 | 0.598 | si:ch73-281n10.2 | -0.366 |
| mbd5 | 0.581 | hnrnpub | -0.359 |
| man2c1 | 0.576 | ppig | -0.357 |
| ccdc171 | 0.573 | NC-002333.4 | -0.353 |
| fam189a1 | 0.564 | anp32e | -0.352 |
| nms | 0.560 | ncl | -0.351 |
| rhbdd1 | 0.560 | srrm1 | -0.346 |
| bahcc1b | 0.559 | hnrnpa1b | -0.341 |
| slf1 | 0.544 | marcksb | -0.338 |
| BX957317.2 | 0.538 | akap12b | -0.334 |
| LOC108183900 | 0.536 | ilf3b | -0.330 |
| stap2b | 0.533 | h1m | -0.328 |
| crfb1 | 0.526 | syncrip | -0.328 |
| zgc:66484 | 0.524 | ptmab | -0.315 |
| FP102052.2 | 0.524 | nucks1a | -0.312 |
| LOC103911261 | 0.523 | hnrnpabb | -0.310 |
| zak | 0.522 | fbl | -0.307 |
| klf7a | 0.518 | stm | -0.306 |
| dhrs11a | 0.517 | epcam | -0.301 |
| myl13 | 0.513 | brd3a | -0.296 |
| prozb | 0.510 | ccna2 | -0.292 |
| mto1 | 0.508 | cd2bp2 | -0.292 |
| appl2 | 0.505 | elavl1a | -0.291 |
| zgc:194551 | 0.503 | hnrnpa1a | -0.290 |
| edil3a | 0.501 | srrt | -0.286 |
| NPAS3 | 0.501 | apoeb | -0.283 |
| zgc:65895 | 0.501 | setb | -0.283 |
| si:ch1073-358c10.1 | 0.499 | tbx16 | -0.281 |
| BX119963.2 | 0.499 | npm1a | -0.280 |
| map6b | 0.494 | nasp | -0.277 |
| itm2bb | 0.494 | hdgfl2 | -0.275 |
| si:ch211-269e2.1 | 0.492 | anp32b | -0.274 |
| cutc | 0.492 | smc2 | -0.274 |
| UNC13A | 0.491 | banf1 | -0.272 |