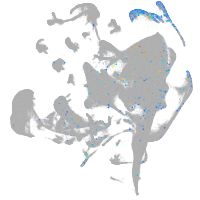
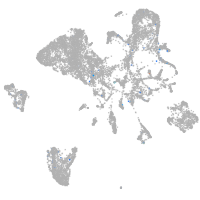
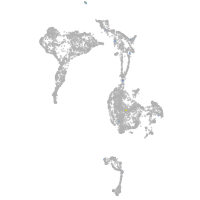
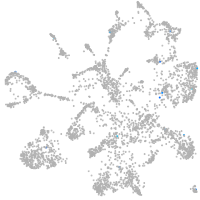
ankyrin repeat and SOCS box containing 15a
ZFIN 
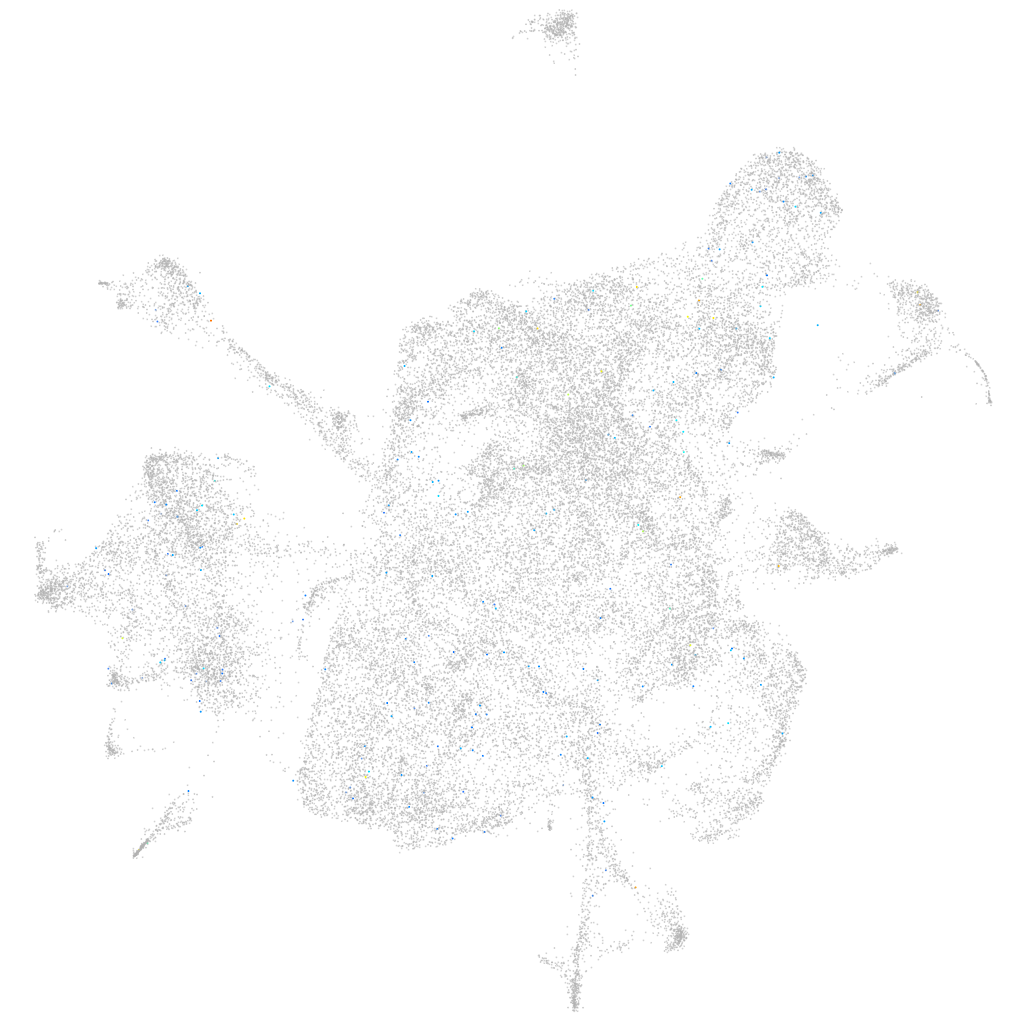
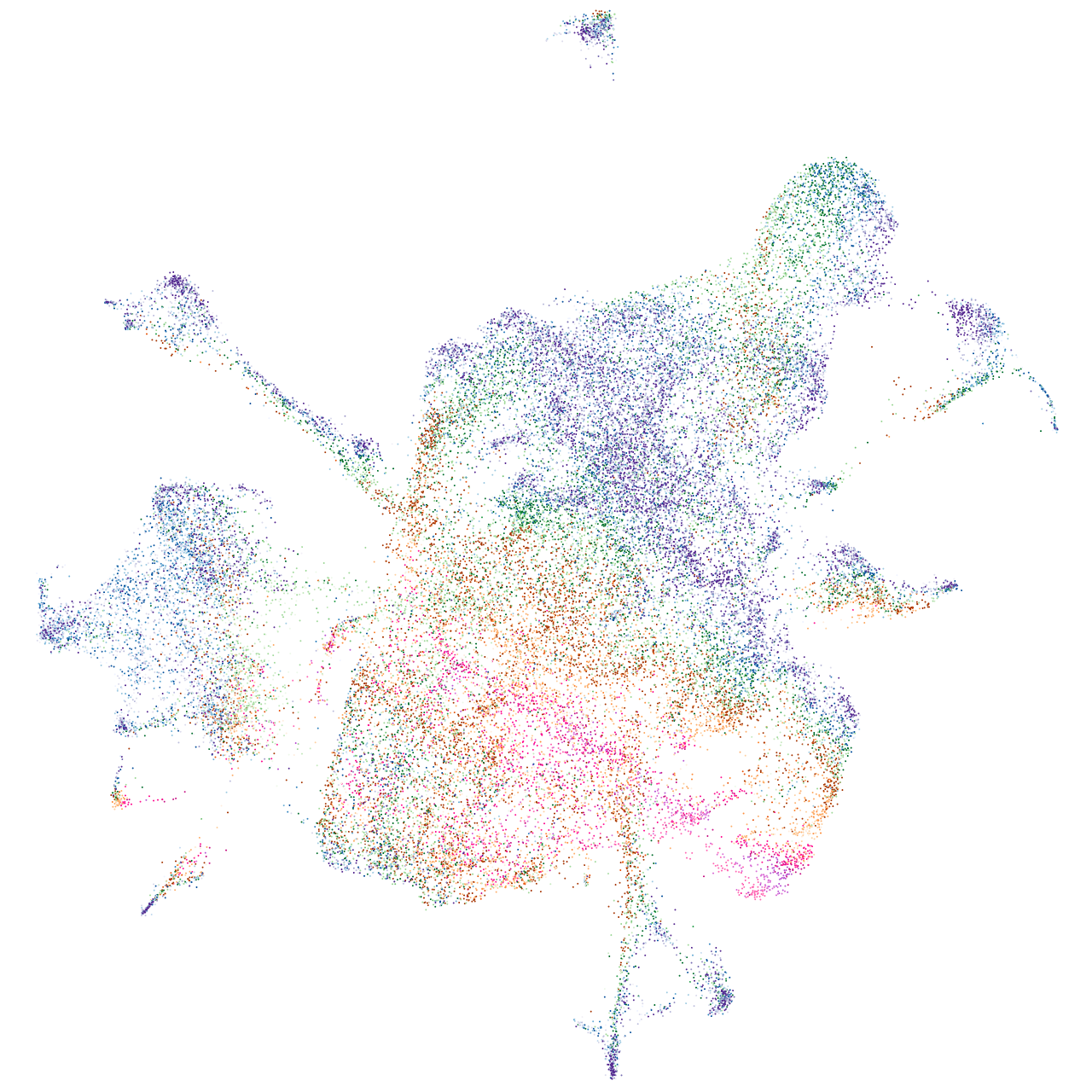
Expression by stage/cluster


Correlated gene expression