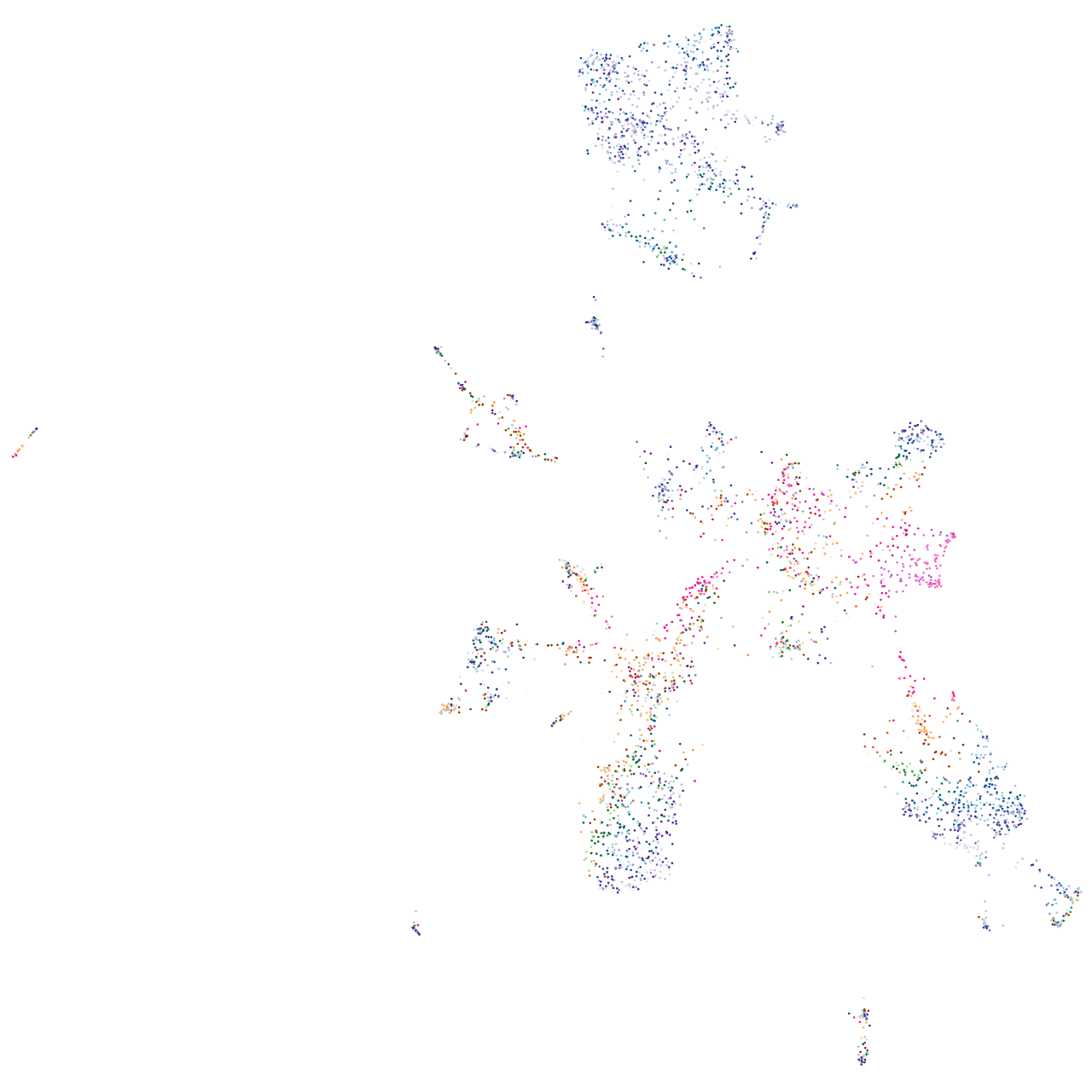
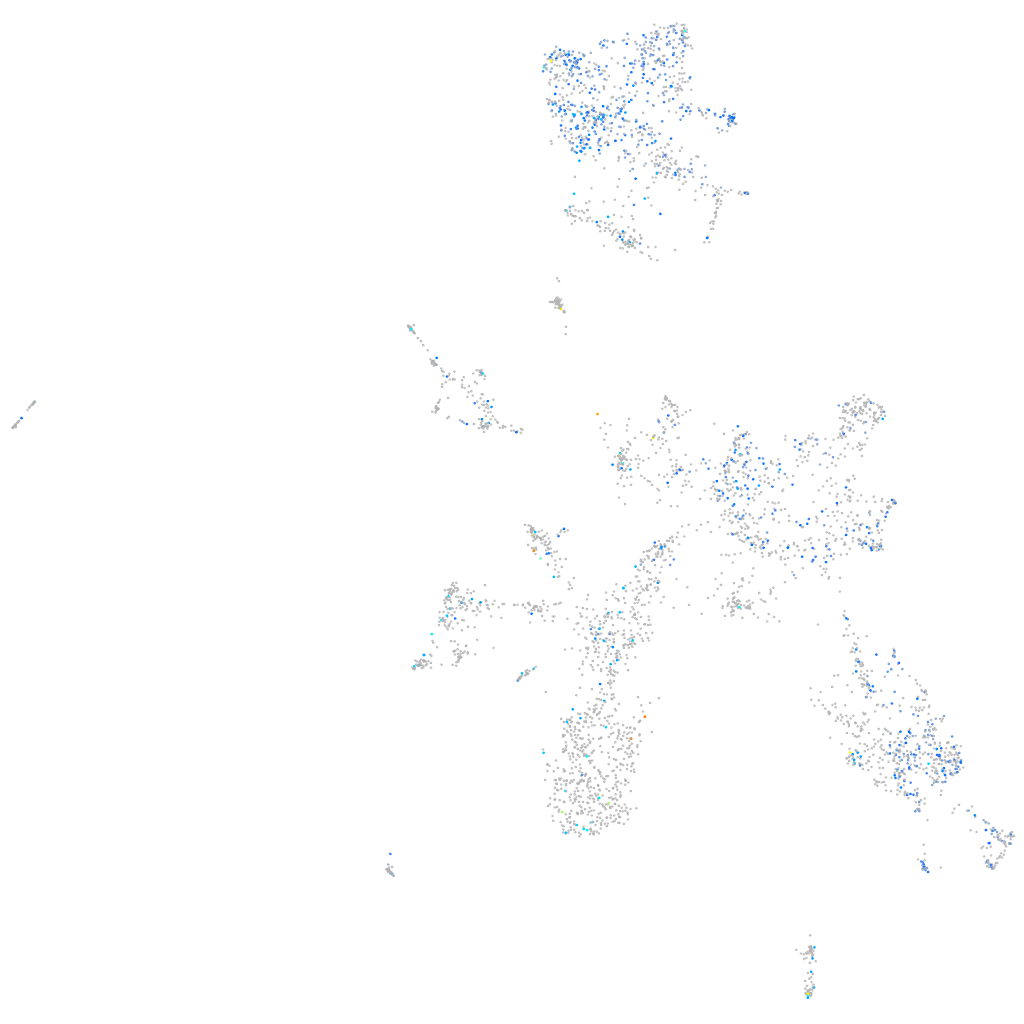
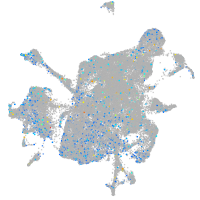
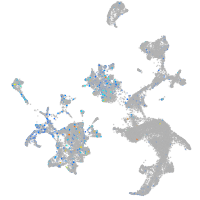
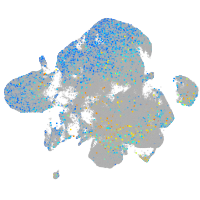
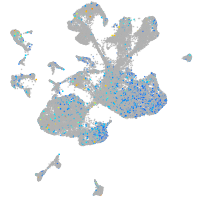
Rho GTPase activating protein 21b
ZFIN 
Expression by stage/cluster


Correlated gene expression