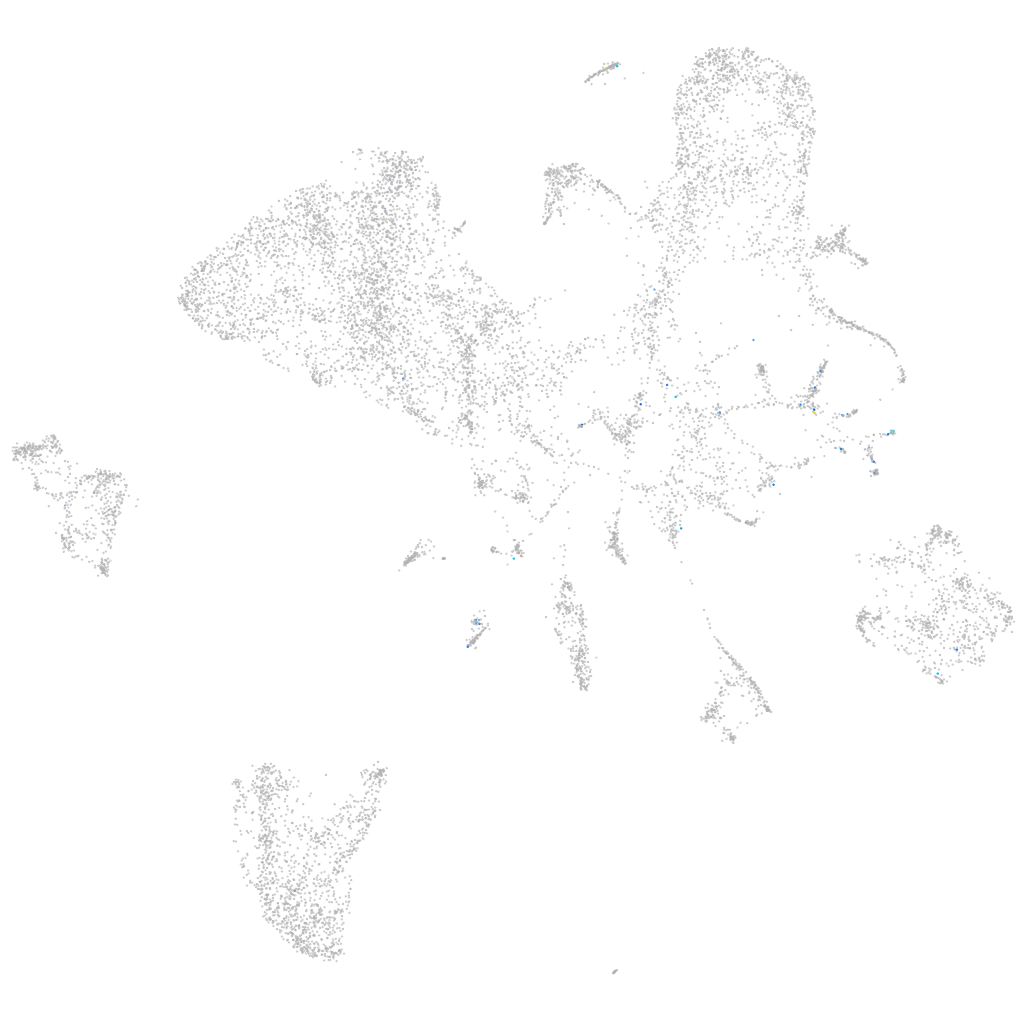
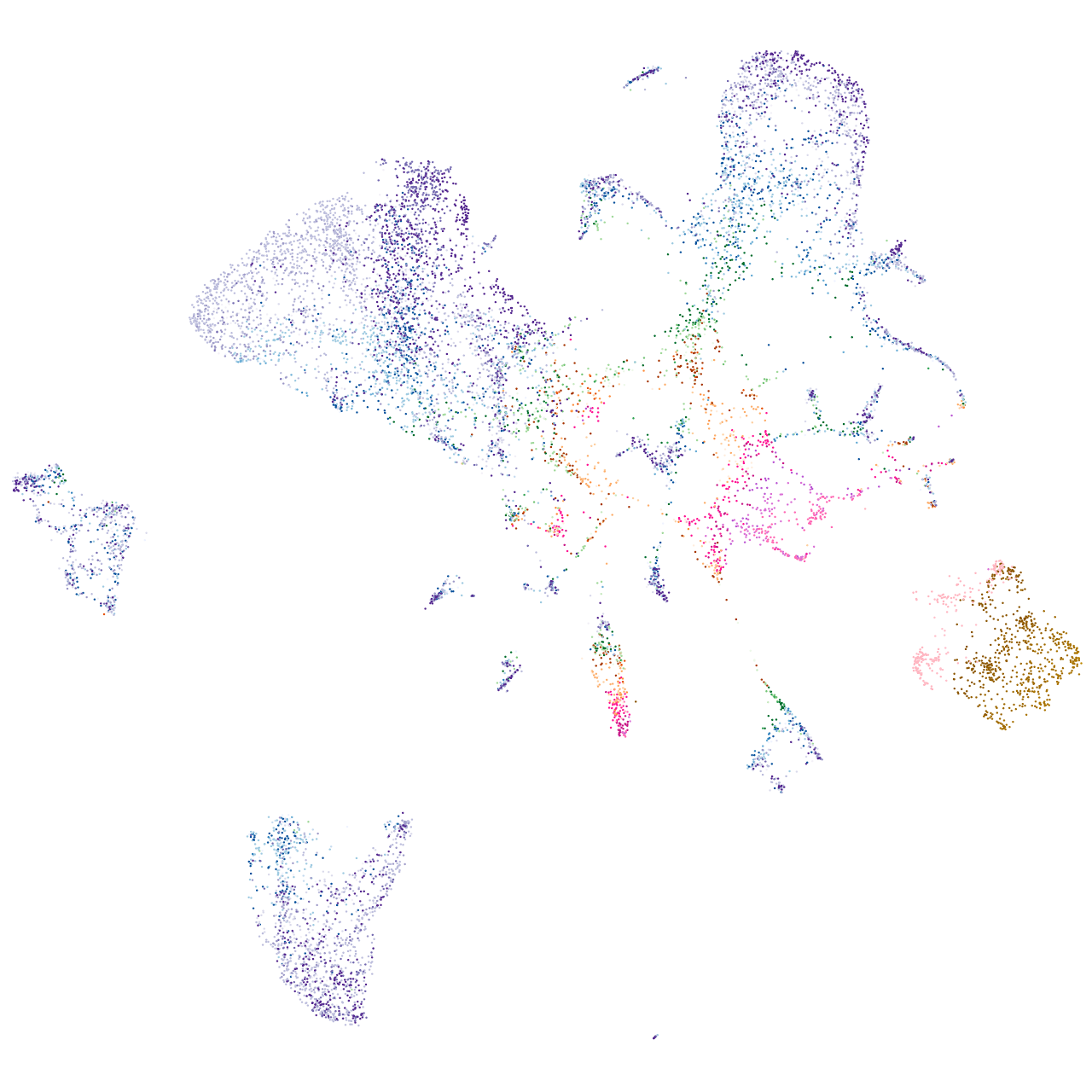
adenylate cyclase 5
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| CR354372.1 | 0.304 | ahcy | -0.056 |
| ptger1b | 0.262 | aldob | -0.055 |
| CABZ01015815.2 | 0.252 | gapdh | -0.050 |
| zbbx | 0.247 | gamt | -0.049 |
| dnah2 | 0.245 | gstp1 | -0.044 |
| fam155a | 0.239 | eno3 | -0.044 |
| XLOC-008708 | 0.229 | fbp1b | -0.044 |
| si:dkey-43k4.5 | 0.226 | gatm | -0.043 |
| hdhd5 | 0.224 | apoa4b.1 | -0.043 |
| CABZ01068208.1 | 0.218 | apoc2 | -0.042 |
| LOC110437784 | 0.216 | bhmt | -0.041 |
| CR339041.1 | 0.210 | glud1b | -0.041 |
| pimr67 | 0.210 | apoa1b | -0.041 |
| si:dkey-65b13.13 | 0.209 | aldh6a1 | -0.040 |
| trim35-30 | 0.207 | afp4 | -0.040 |
| ptprja | 0.207 | gstt1a | -0.039 |
| LOC100330246 | 0.204 | mat1a | -0.038 |
| gabrb1 | 0.197 | scp2a | -0.038 |
| LOC101883236 | 0.177 | nupr1b | -0.038 |
| pmfbp1 | 0.175 | gpx4a | -0.038 |
| BX649476.1 | 0.170 | cx32.3 | -0.038 |
| lingo2a | 0.169 | acadm | -0.038 |
| casr | 0.166 | dap | -0.038 |
| si:ch73-233f7.7 | 0.162 | sult2st2 | -0.036 |
| irf4a | 0.155 | ugt1a7 | -0.036 |
| si:ch211-214b16.4 | 0.155 | gcshb | -0.036 |
| si:ch73-249k16.4 | 0.155 | ppa1b | -0.036 |
| diras2 | 0.153 | sdr16c5b | -0.036 |
| sgip1b | 0.145 | gnmt | -0.035 |
| LOC100537404 | 0.143 | agxtb | -0.035 |
| CT573316.3 | 0.142 | cbr1l | -0.035 |
| neurod1 | 0.141 | g6pca.2 | -0.035 |
| ghrl | 0.140 | apobb.1 | -0.035 |
| vwa5b2 | 0.140 | suclg1 | -0.035 |
| adamts5 | 0.139 | nipsnap3a | -0.034 |