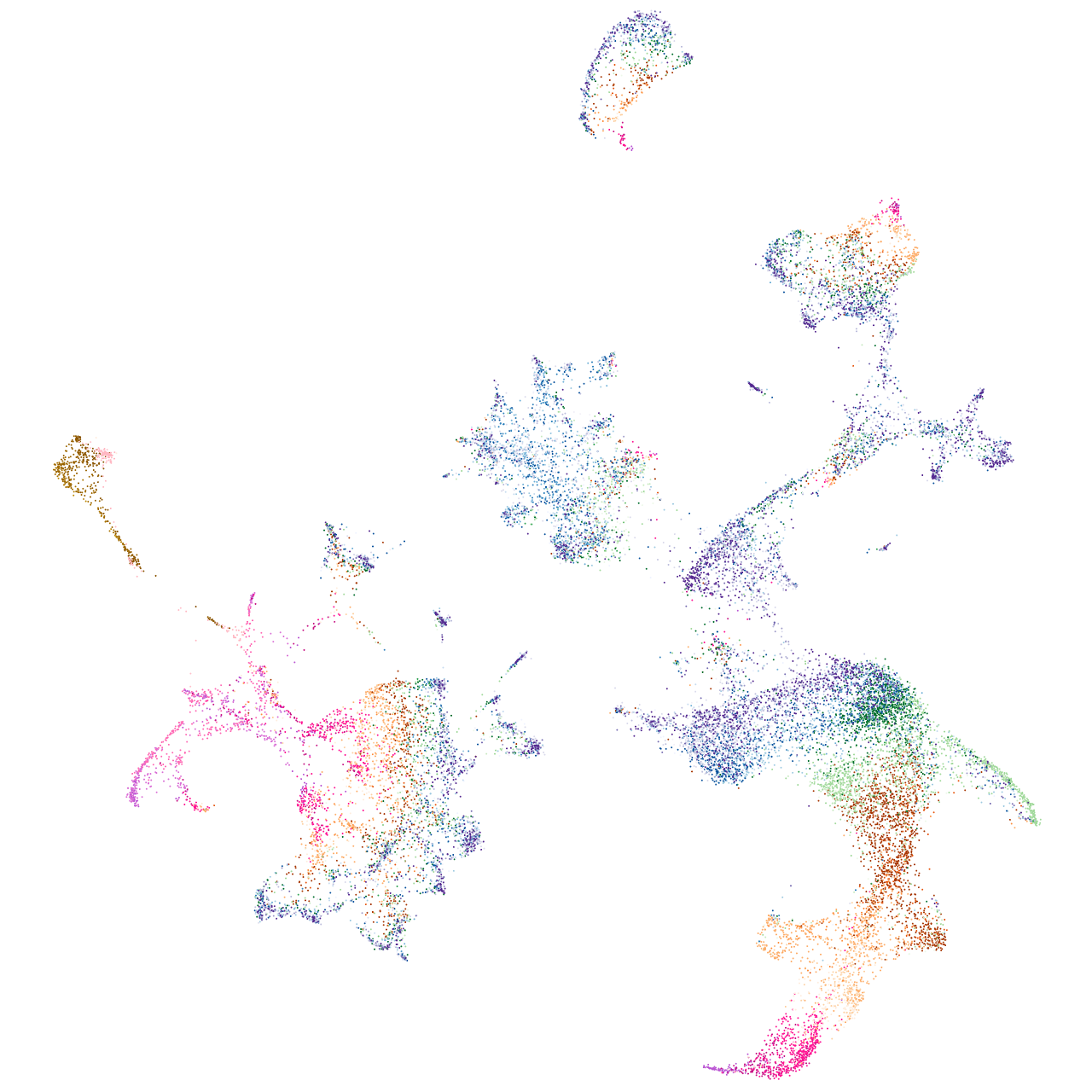zgc:91968
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| opn5 | 0.213 | metap2b | -0.021 |
| tyrp1b | 0.209 | minos1 | -0.017 |
| mlpha | 0.194 | selenok | -0.014 |
| ankrd2 | 0.187 | cox4i1 | -0.014 |
| si:ch73-389b16.1 | 0.158 | ywhaqa | -0.014 |
| tyrp1a | 0.156 | psmb2 | -0.014 |
| kcnk12l | 0.149 | elavl1a | -0.013 |
| pmela | 0.148 | anxa11a | -0.013 |
| cbln2a | 0.144 | CU694197.1 | -0.013 |
| CR388166.1 | 0.127 | ano6 | -0.013 |
| opn7d | 0.127 | si:ch211-284b7.3 | -0.013 |
| pthlha | 0.122 | zgc:86598 | -0.013 |
| oca2 | 0.119 | ppp6r3 | -0.013 |
| CU462913.1 | 0.117 | tram1 | -0.013 |
| gls2b | 0.112 | serbp1a | -0.013 |
| XLOC-021942 | 0.111 | qars | -0.013 |
| krt93 | 0.110 | rfc3 | -0.013 |
| LECT2 (1 of many) | 0.109 | atp5f1d | -0.013 |
| dct | 0.107 | wdr45b | -0.013 |
| plppr5a | 0.106 | nop56 | -0.013 |
| slc45a2 | 0.106 | immt | -0.013 |
| rsph14 | 0.101 | uchl5 | -0.013 |
| F7 (1 of many) | 0.100 | zgc:153409 | -0.013 |
| kcnk2b | 0.099 | cox17 | -0.012 |
| LOC101882691 | 0.099 | pam16 | -0.012 |
| XLOC-019062 | 0.098 | ndufs3 | -0.012 |
| BX547928.1 | 0.098 | eef1da | -0.012 |
| smtna | 0.096 | atp5pf | -0.012 |
| cnga4 | 0.096 | zfand5b | -0.012 |
| LOC103909610 | 0.095 | rps6kb1b | -0.012 |
| ido1 | 0.090 | tet3 | -0.012 |
| th2 | 0.090 | fam107b | -0.012 |
| kcnk17 | 0.090 | tmem256 | -0.012 |
| CABZ01044235.1 | 0.089 | myd88 | -0.012 |
| gpr143 | 0.086 | gmfg | -0.012 |