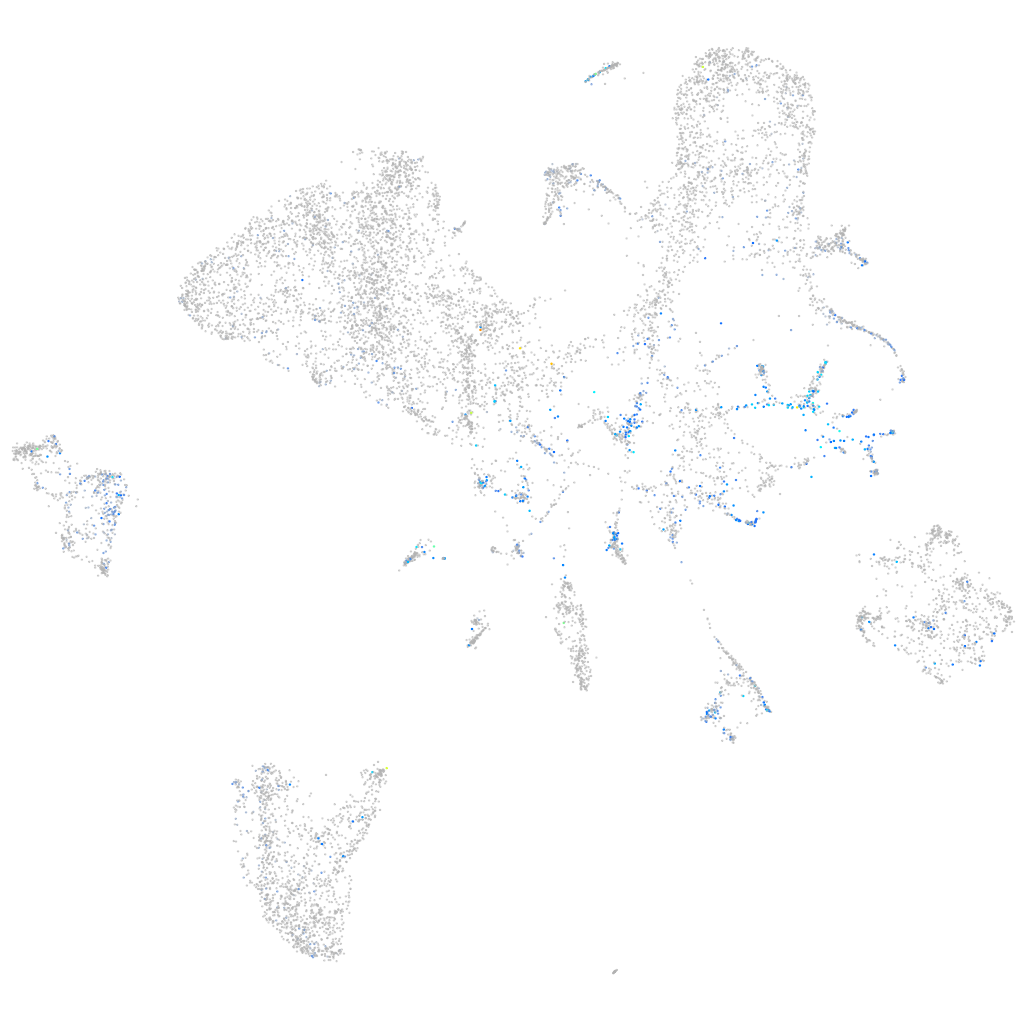
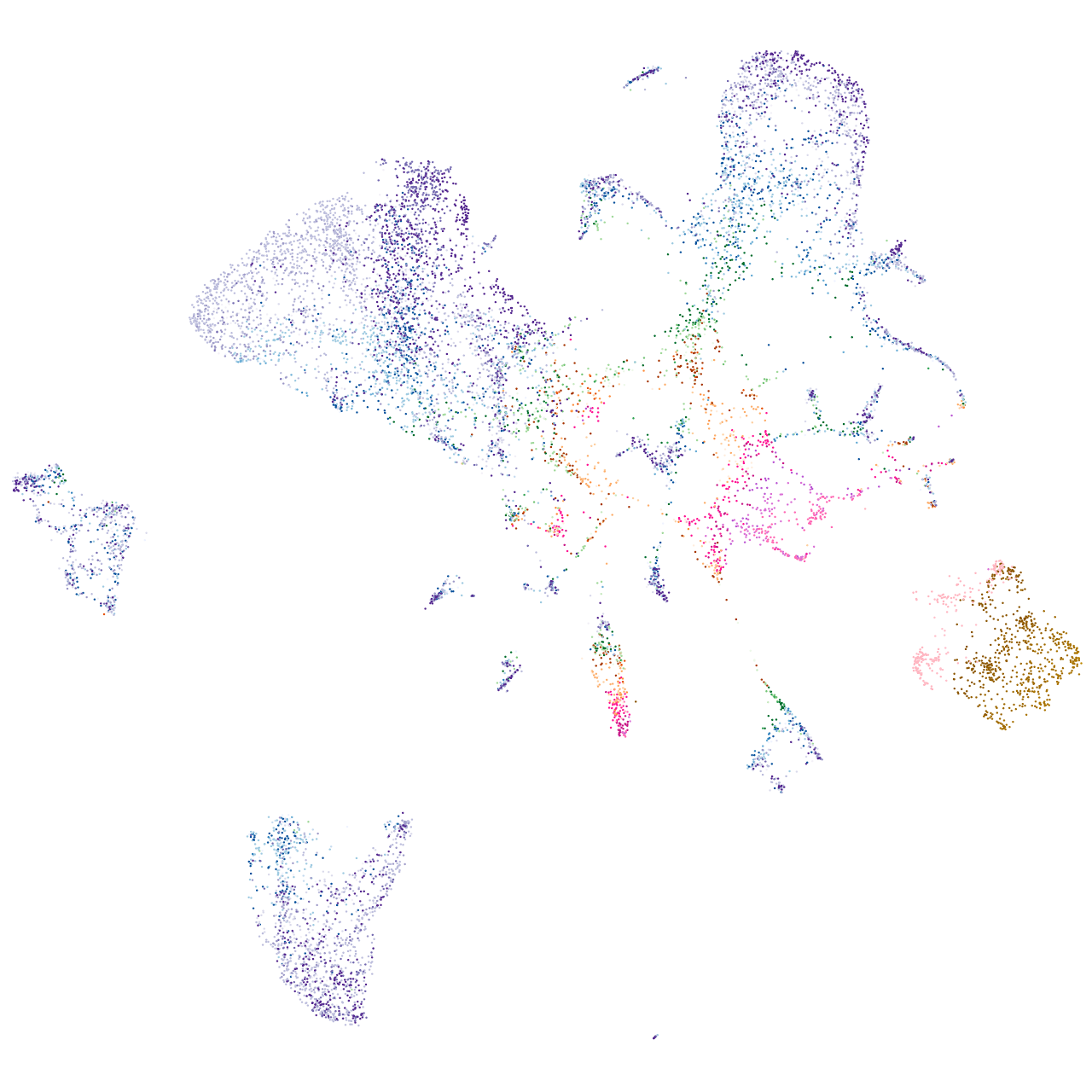
TOX high mobility group box family member 3
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| insm1a | 0.324 | gamt | -0.191 |
| neurod1 | 0.276 | gapdh | -0.185 |
| scg3 | 0.259 | ahcy | -0.169 |
| dlb | 0.246 | apoa4b.1 | -0.166 |
| tox | 0.245 | aldob | -0.163 |
| mir375-2 | 0.234 | gatm | -0.163 |
| insm1b | 0.231 | fbp1b | -0.162 |
| dll4 | 0.230 | apoc2 | -0.161 |
| myt1b | 0.226 | apoa1b | -0.160 |
| elovl1a | 0.225 | glud1b | -0.159 |
| mir7a-1 | 0.224 | scp2a | -0.153 |
| arxa | 0.222 | bhmt | -0.153 |
| nkx2.2a | 0.221 | mat1a | -0.152 |
| jpt1b | 0.217 | afp4 | -0.151 |
| si:dkey-153k10.9 | 0.213 | gcshb | -0.145 |
| ret | 0.212 | gpx4a | -0.144 |
| id4 | 0.210 | gstt1a | -0.143 |
| sox4b | 0.209 | agxtb | -0.143 |
| isl1 | 0.209 | cx32.3 | -0.141 |
| si:ch1073-429i10.3.1 | 0.208 | pnp4b | -0.141 |
| tmsb | 0.208 | rdh1 | -0.140 |
| kdm6bb | 0.206 | apobb.1 | -0.139 |
| si:dkey-42i9.4 | 0.205 | mgst1.2 | -0.139 |
| scgn | 0.205 | apoc1 | -0.138 |
| vamp2 | 0.204 | suclg2 | -0.138 |
| lysmd2 | 0.202 | aldh6a1 | -0.136 |
| anxa4 | 0.200 | tdo2a | -0.136 |
| soga1 | 0.200 | apoa2 | -0.135 |
| pax6b | 0.196 | etnppl | -0.135 |
| ywhah | 0.194 | abat | -0.135 |
| hepacam2 | 0.194 | slco1d1 | -0.134 |
| c2cd4a | 0.193 | aldh7a1 | -0.134 |
| si:dkey-28b4.7 | 0.190 | dhrs9 | -0.133 |
| hmgb1a | 0.188 | nipsnap3a | -0.133 |
| oaz1b | 0.188 | apom | -0.132 |