si:ch73-97h19.2
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs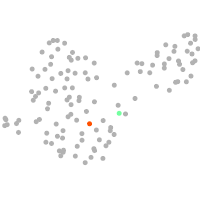 |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| LOC101884422 | 0.459 | gapdh | -0.034 |
| si:dkey-266f7.4 | 0.343 | gamt | -0.034 |
| LOC100148648 | 0.334 | dbi | -0.031 |
| LOC110440093 | 0.306 | hspe1 | -0.031 |
| si:ch211-255a21.2 | 0.252 | gpx4a | -0.031 |
| FO834800.1 | 0.245 | fbp1b | -0.030 |
| lipib | 0.229 | lgals2b | -0.029 |
| CR388231.2 | 0.229 | aldh9a1a.1 | -0.029 |
| LOC103909738 | 0.225 | pklr | -0.028 |
| CABZ01072815.1 | 0.222 | slc25a20 | -0.028 |
| rnf217 | 0.218 | gpx1a | -0.027 |
| slc1a7a | 0.216 | gatm | -0.027 |
| LOC569262 | 0.215 | suclg1 | -0.027 |
| vwa2 | 0.206 | eno3 | -0.027 |
| ch25hl2 | 0.193 | cox6b1 | -0.026 |
| LOC101884267 | 0.184 | mat1a | -0.026 |
| slc6a2 | 0.169 | gstr | -0.026 |
| ppp2r2cb | 0.166 | apoa4b.1 | -0.026 |
| lhx8a | 0.164 | glud1b | -0.025 |
| trim35-31 | 0.161 | sdr16c5b | -0.025 |
| trim35-9 | 0.155 | aldh7a1 | -0.025 |
| scel | 0.153 | gcshb | -0.025 |
| ftr38 | 0.153 | BX908782.3 | -0.025 |
| soga3b | 0.151 | scp2a | -0.025 |
| mamdc2b | 0.149 | nipsnap3a | -0.025 |
| si:zfos-2330d3.1 | 0.142 | grhprb | -0.024 |
| creb5b | 0.141 | sult2st2 | -0.024 |
| syt7a | 0.140 | dhdhl | -0.024 |
| LOC100331080 | 0.135 | gstt1a | -0.024 |
| si:ch211-157c3.4 | 0.134 | ahcy | -0.023 |
| krt1-19d | 0.133 | abat | -0.023 |
| si:dkeyp-51f12.2 | 0.132 | afp4 | -0.023 |
| adgrf6 | 0.131 | cox7a1 | -0.023 |
| svopa | 0.130 | fdx1 | -0.023 |
| ponzr2 | 0.127 | apoc2 | -0.023 |