si:ch211-195b15.7
ZFIN 
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line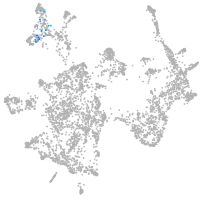 |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| CU466278.2 | 0.566 | eef1a1l1 | -0.086 |
| XLOC-041698 | 0.494 | eef2b | -0.073 |
| foxp3a | 0.411 | rps11 | -0.072 |
| BX294379.1 | 0.330 | naca | -0.071 |
| BX276107.1 | 0.260 | rpl32 | -0.070 |
| LOC110438172 | 0.245 | rpl27 | -0.069 |
| LOC101885030 | 0.197 | h3f3d | -0.069 |
| amfrb | 0.196 | serbp1a | -0.068 |
| prune2 | 0.192 | rpl7a | -0.067 |
| PLEKHB1 | 0.188 | rplp1 | -0.067 |
| CABZ01117780.1 | 0.179 | hspa8 | -0.067 |
| ldhbb | 0.173 | rpl13a | -0.067 |
| cyp2ad2 | 0.168 | rps9 | -0.066 |
| rom1a | 0.161 | cirbpb | -0.066 |
| gls2b | 0.161 | rps19 | -0.066 |
| adamts15b | 0.161 | tpt1 | -0.065 |
| sstr2a | 0.158 | hsp90ab1 | -0.063 |
| slc6a17 | 0.157 | rpl36 | -0.063 |
| saga | 0.156 | rpsa | -0.063 |
| ccdc135 | 0.153 | rps16 | -0.063 |
| snx19b | 0.148 | rpl7 | -0.062 |
| fbxl17 | 0.146 | rack1 | -0.062 |
| zgc:171482 | 0.141 | rps15 | -0.062 |
| guca1b | 0.141 | rps12 | -0.062 |
| cabp4 | 0.137 | pabpc1a | -0.061 |
| asb14b | 0.132 | rpl26 | -0.061 |
| loxl5a | 0.130 | rpl13 | -0.061 |
| cnga1b | 0.129 | rps14 | -0.060 |
| si:ch211-113d22.2 | 0.126 | h2afvb | -0.060 |
| gnat1 | 0.125 | rplp2l | -0.060 |
| rom1b | 0.124 | rpl14 | -0.060 |
| fabp6 | 0.119 | ran | -0.060 |
| sagb | 0.110 | khdrbs1a | -0.059 |
| atp11a | 0.106 | rps3a | -0.058 |
| asb2a.1 | 0.102 | rpl35 | -0.058 |