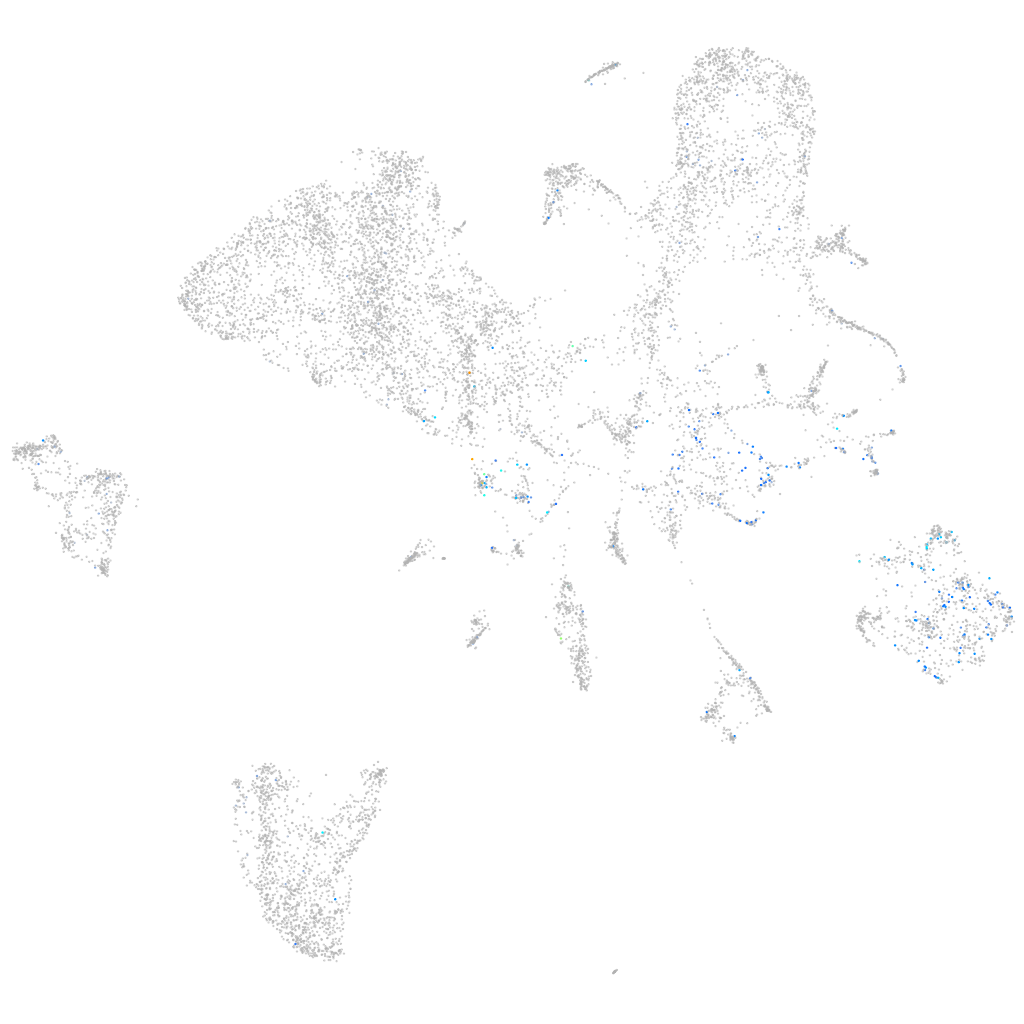
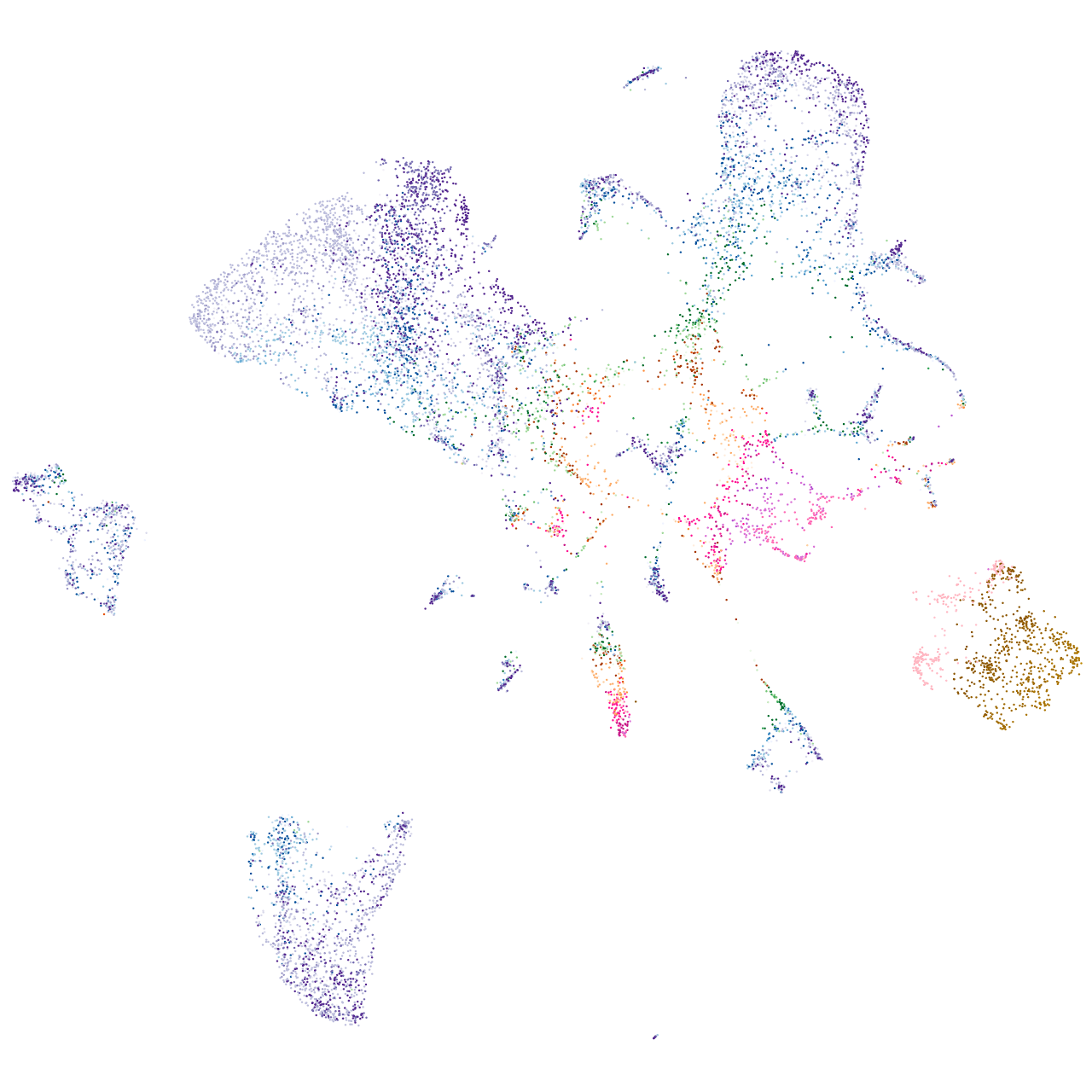
secretory carrier membrane protein 5a
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| marcksl1b | 0.187 | nme2b.1 | -0.126 |
| marcksb | 0.183 | ahcy | -0.122 |
| cx43.4 | 0.181 | rps10 | -0.120 |
| tulp4b | 0.179 | gapdh | -0.120 |
| dicp1.5-6 | 0.175 | eno3 | -0.116 |
| fcer1g | 0.174 | rpl37 | -0.114 |
| myf6 | 0.164 | gamt | -0.113 |
| cdh2 | 0.163 | sod1 | -0.111 |
| cdx4 | 0.161 | zgc:114188 | -0.111 |
| hspb1 | 0.160 | eef1da | -0.111 |
| nucks1a | 0.160 | rps17 | -0.106 |
| cacng8b | 0.159 | suclg1 | -0.104 |
| anp32e | 0.158 | aldob | -0.103 |
| si:dkey-40m6.8 | 0.157 | nupr1b | -0.101 |
| ss18 | 0.157 | glud1b | -0.101 |
| il17rd | 0.155 | fbp1b | -0.100 |
| hp1bp3 | 0.155 | gstp1 | -0.100 |
| hmgb1b | 0.155 | aldh7a1 | -0.098 |
| znfl2a | 0.155 | gatm | -0.097 |
| akap12b | 0.154 | prdx2 | -0.094 |
| dnmt3bb.2 | 0.153 | BX908782.3 | -0.093 |
| akap6 | 0.153 | pgk1 | -0.093 |
| ncf2 | 0.152 | atp5if1b | -0.092 |
| si:ch211-137a8.4 | 0.152 | sdr16c5b | -0.091 |
| zgc:110425 | 0.152 | atp5pf | -0.091 |
| h3f3d | 0.152 | gpx4a | -0.091 |
| cd99l2 | 0.151 | scp2a | -0.091 |
| hnrnpa0a | 0.151 | gstt1a | -0.091 |
| khdrbs1a | 0.150 | cx32.3 | -0.091 |
| hmga1a | 0.150 | ckba | -0.090 |
| syncrip | 0.150 | mat1a | -0.090 |
| ilf3b | 0.148 | apoa4b.1 | -0.089 |
| anp32b | 0.148 | ddt | -0.089 |
| gper1 | 0.148 | atp5fa1 | -0.089 |
| hnrnpaba | 0.147 | rmdn1 | -0.088 |