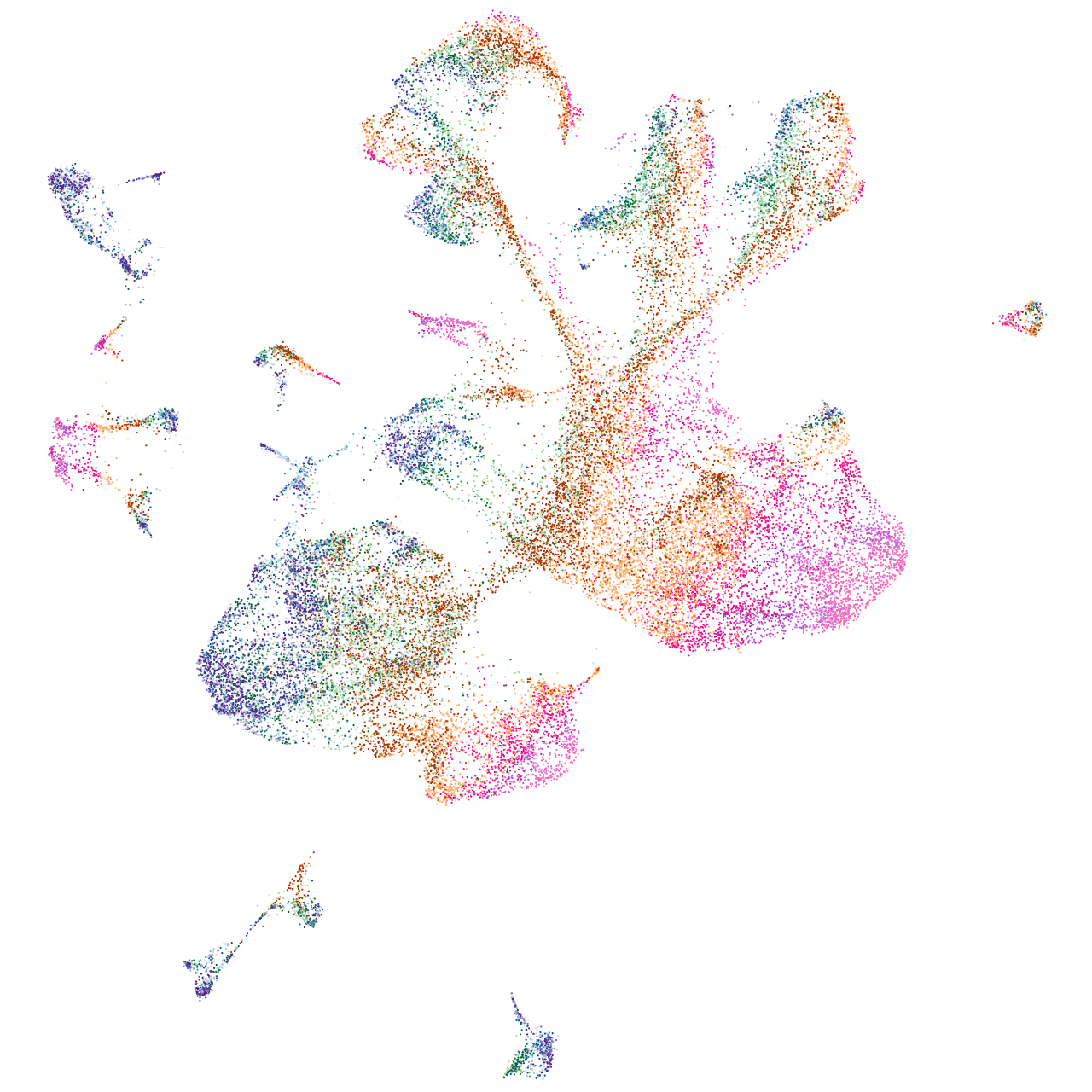
ribosomal protein S5
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| rps7 | 0.653 | glula | -0.369 |
| rps20 | 0.646 | CR383676.1 | -0.367 |
| rps23 | 0.630 | acbd7 | -0.358 |
| rpl23 | 0.626 | cx43 | -0.342 |
| rplp0 | 0.626 | gapdhs | -0.341 |
| rplp1 | 0.625 | slc1a2b | -0.336 |
| rpl7a | 0.625 | ckbb | -0.335 |
| rplp2l | 0.625 | aldocb | -0.334 |
| rps3a | 0.624 | mt2 | -0.324 |
| rpl7 | 0.624 | efhd1 | -0.314 |
| rpl10a | 0.622 | slc4a4a | -0.310 |
| rps9 | 0.621 | slc6a1b | -0.305 |
| rpl13 | 0.621 | ndrg3a | -0.303 |
| rpl21 | 0.621 | IGLON5 | -0.279 |
| rps12 | 0.620 | gpm6ab | -0.278 |
| rpl8 | 0.620 | gpm6aa | -0.274 |
| rpl11 | 0.618 | gpr37l1b | -0.274 |
| rps19 | 0.617 | elavl4 | -0.273 |
| rps15a | 0.617 | sncgb | -0.272 |
| rpl26 | 0.614 | slc38a3a | -0.267 |
| rpl10 | 0.614 | cdo1 | -0.266 |
| rps14 | 0.613 | ppap2d | -0.266 |
| rps27a | 0.612 | cadm4 | -0.266 |
| rpl12 | 0.611 | smox | -0.265 |
| rpl19 | 0.608 | slc3a2a | -0.264 |
| rps4x | 0.607 | cyp2ad3 | -0.259 |
| rpl17 | 0.604 | ptn | -0.258 |
| rps2 | 0.604 | pygmb | -0.257 |
| rpsa | 0.603 | mfge8a | -0.256 |
| rps27.1 | 0.602 | rnasekb | -0.256 |
| si:dkey-151g10.6 | 0.600 | atp6v0cb | -0.255 |
| rps13 | 0.599 | spock2 | -0.254 |
| rpl28 | 0.598 | ak1 | -0.251 |
| rps10 | 0.598 | pvalb1 | -0.250 |
| rps24 | 0.597 | cahz | -0.249 |