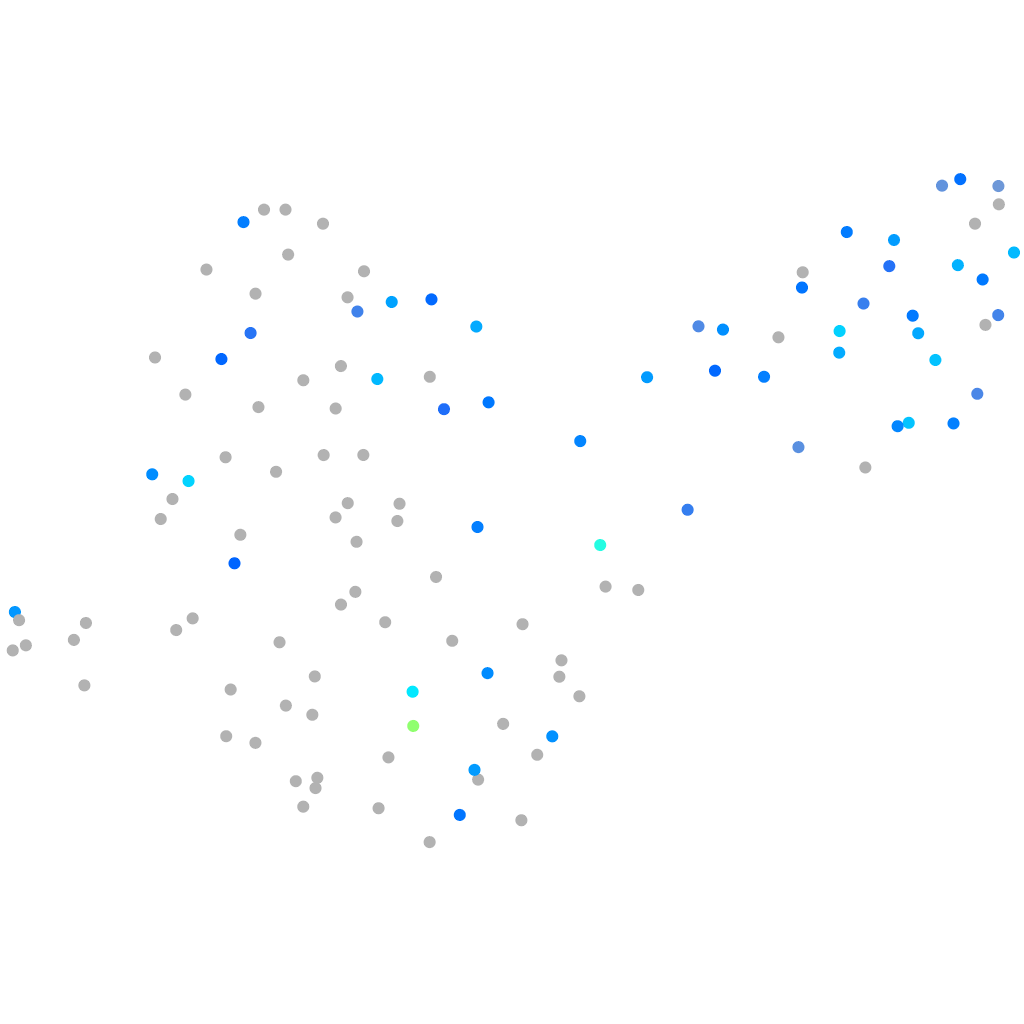
mitochondrial pyruvate carrier 2
ZFIN 
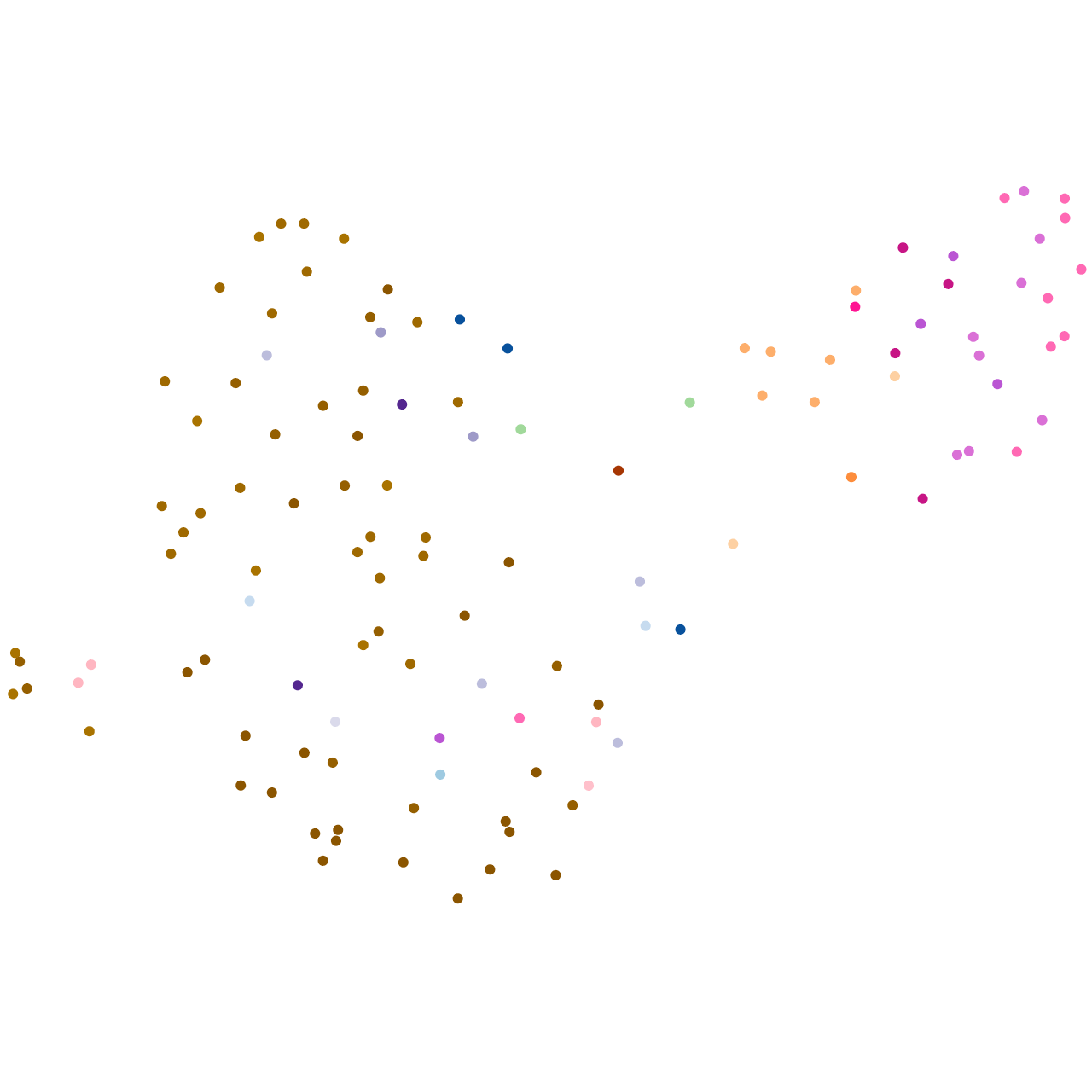
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| dap1b | 0.586 | ncl | -0.553 |
| prdx5 | 0.583 | NC-002333.4 | -0.526 |
| zgc:56493 | 0.558 | hnrnpa1b | -0.496 |
| rpl37 | 0.534 | acin1b | -0.483 |
| CABZ01072614.1 | 0.531 | ppig | -0.454 |
| zgc:114188 | 0.527 | tbx16 | -0.440 |
| rnasekb | 0.523 | h1m | -0.436 |
| rps10 | 0.521 | snrpa | -0.429 |
| cox5ab | 0.514 | akap12b | -0.428 |
| rps27.1 | 0.512 | hnrnpub | -0.426 |
| eif3k | 0.511 | u2surp | -0.416 |
| tmed4 | 0.509 | fbl | -0.415 |
| rps17 | 0.507 | slu7 | -0.413 |
| tnrc6a | 0.504 | tpx2 | -0.413 |
| zgc:86839 | 0.502 | stmn1a | -0.412 |
| ndufb7 | 0.501 | hnrnpa1a | -0.411 |
| mycbp | 0.501 | ved | -0.411 |
| cct2 | 0.500 | marcksb | -0.407 |
| btf3 | 0.498 | nasp | -0.406 |
| slc25a3b | 0.498 | top1l | -0.404 |
| rps9 | 0.496 | ebna1bp2 | -0.403 |
| gabarapb | 0.495 | npm1a | -0.400 |
| cox6a1 | 0.494 | nop56 | -0.398 |
| COX7A2 (1 of many) | 0.492 | syncrip | -0.391 |
| hnrnpa0l | 0.488 | snrpb2 | -0.391 |
| gapdhs | 0.486 | baz1b | -0.382 |
| gabarapl2 | 0.484 | ddit4 | -0.377 |
| rps13 | 0.483 | zgc:112356 | -0.377 |
| ccni | 0.480 | hnrnpd | -0.375 |
| rps24 | 0.472 | stm | -0.371 |
| rpl8 | 0.471 | hnrnpm | -0.370 |
| rps25 | 0.470 | gra | -0.368 |
| pin1 | 0.469 | nucks1b | -0.367 |
| rps14 | 0.469 | brd3a | -0.366 |
| pfdn6 | 0.468 | nop58 | -0.362 |