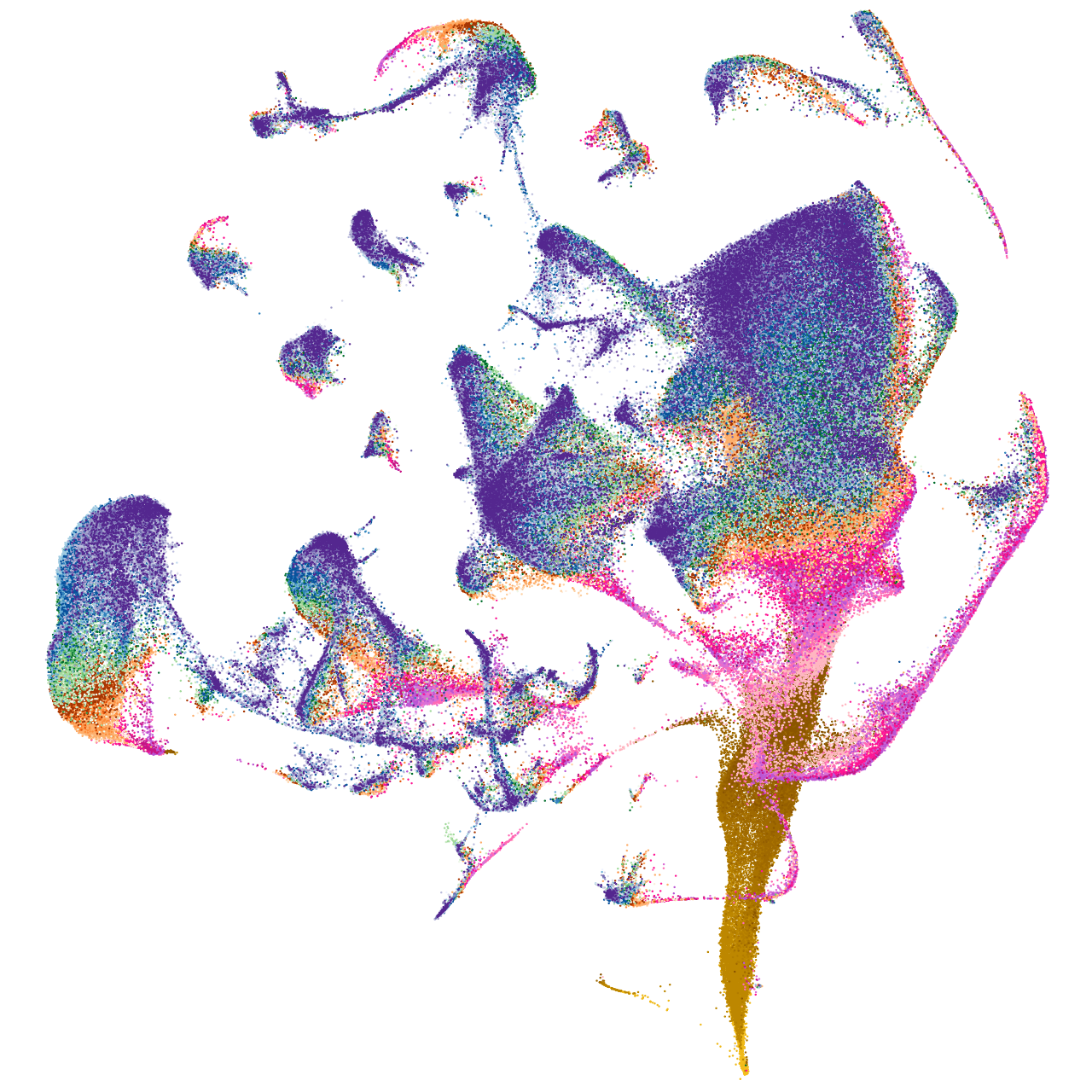Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
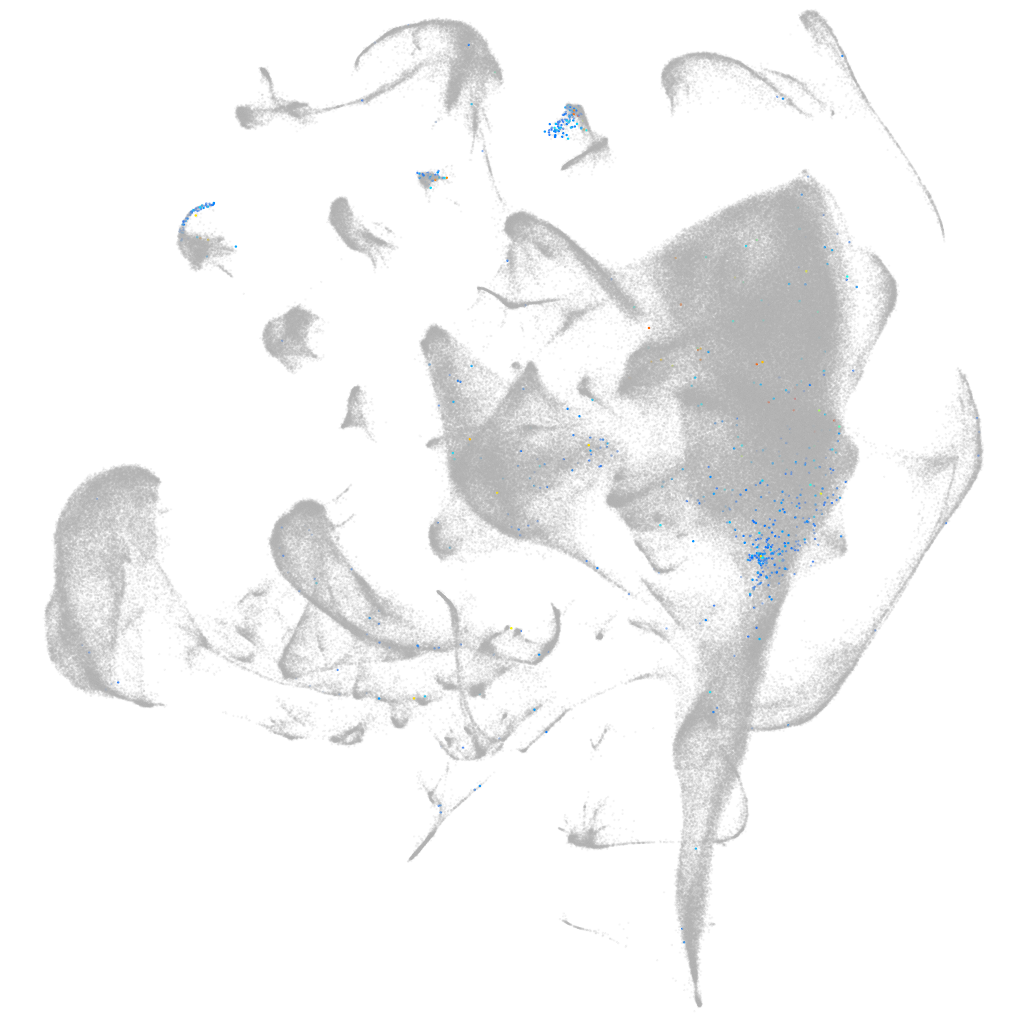
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| crestin | 0.282 | msi1 | -0.015 |
| LOC110438542 | 0.222 | pvalb2 | -0.015 |
| ednrab | 0.205 | col1a2 | -0.014 |
| si:ch211-11c3.9 | 0.132 | nova2 | -0.014 |
| sox10 | 0.123 | pvalb1 | -0.014 |
| mov10b.1 | 0.112 | zgc:162730 | -0.014 |
| tfec | 0.110 | cyt1 | -0.013 |
| trpm1b | 0.108 | krt4 | -0.013 |
| mitfa | 0.104 | actc1b | -0.012 |
| gpr143 | 0.098 | cfl1l | -0.012 |
| CABZ01048402.2 | 0.097 | col1a1a | -0.012 |
| znf1004 | 0.097 | gpm6aa | -0.012 |
| si:ch211-108c6.2 | 0.093 | mdka | -0.012 |
| erbb3b | 0.092 | rbp4 | -0.012 |
| rab38 | 0.088 | tuba1c | -0.012 |
| BX248318.1 | 0.087 | btg2 | -0.011 |
| ednrba | 0.086 | cadm3 | -0.011 |
| rab32a | 0.079 | cldn5a | -0.011 |
| slc45a2 | 0.079 | col1a1b | -0.011 |
| pnp4a | 0.078 | CU467822.1 | -0.011 |
| rab34b | 0.077 | cyt1l | -0.011 |
| emp3b | 0.076 | elavl3 | -0.011 |
| tyr | 0.076 | fgfr2 | -0.011 |
| si:ch73-389b16.1 | 0.074 | krt5 | -0.011 |
| tspan36 | 0.074 | krtt1c19e | -0.011 |
| si:ch211-243a20.3 | 0.074 | mgst1.2 | -0.011 |
| bace2 | 0.073 | pycard | -0.011 |
| slc15a2 | 0.073 | selenow1 | -0.011 |
| smim29 | 0.072 | sox3 | -0.011 |
| cyth1b | 0.071 | tmsb | -0.011 |
| si:ch73-156l6.2 | 0.071 | tmsb1 | -0.011 |
| ly6m5 | 0.070 | tuba8l2 | -0.011 |
| pts | 0.070 | wu:fb18f06 | -0.011 |
| si:dkey-21a6.5 | 0.070 | zgc:165461 | -0.011 |
| FP085398.1 | 0.069 | bhmt | -0.011 |