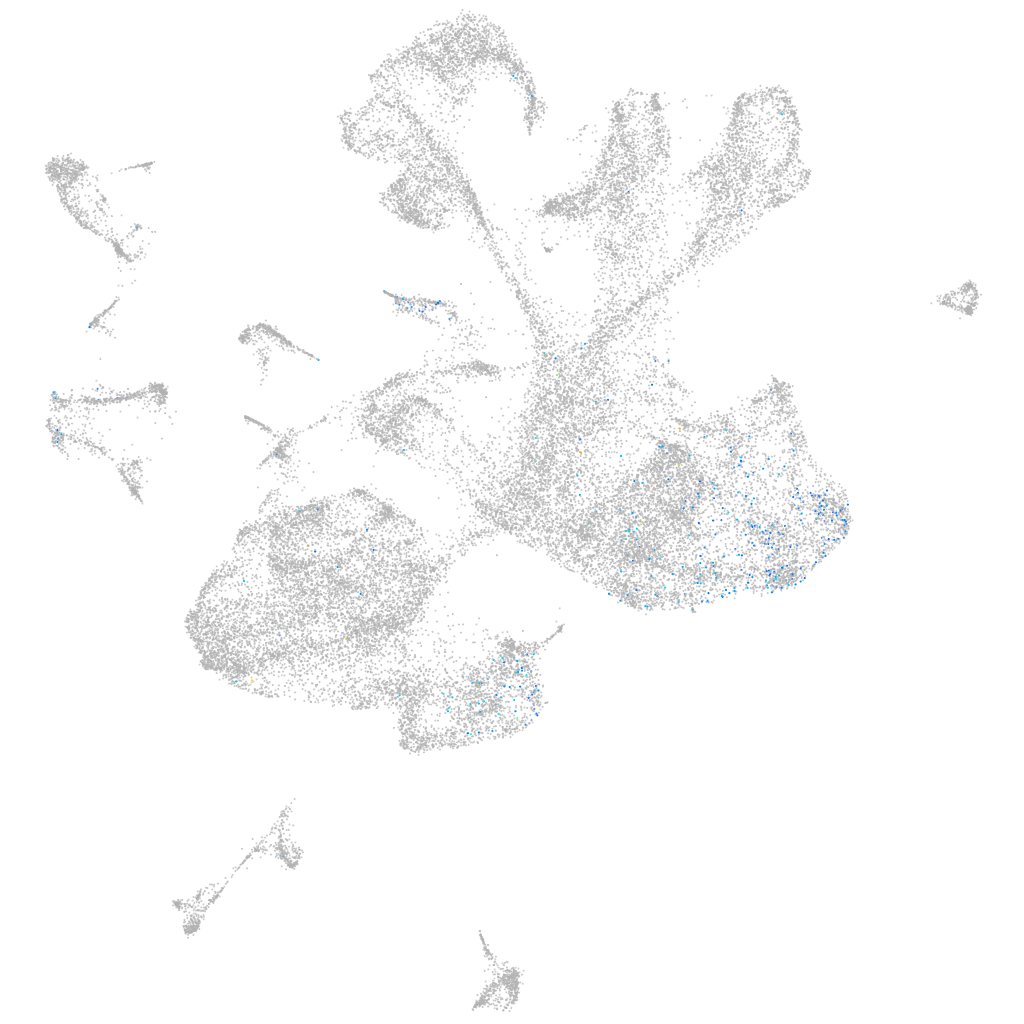
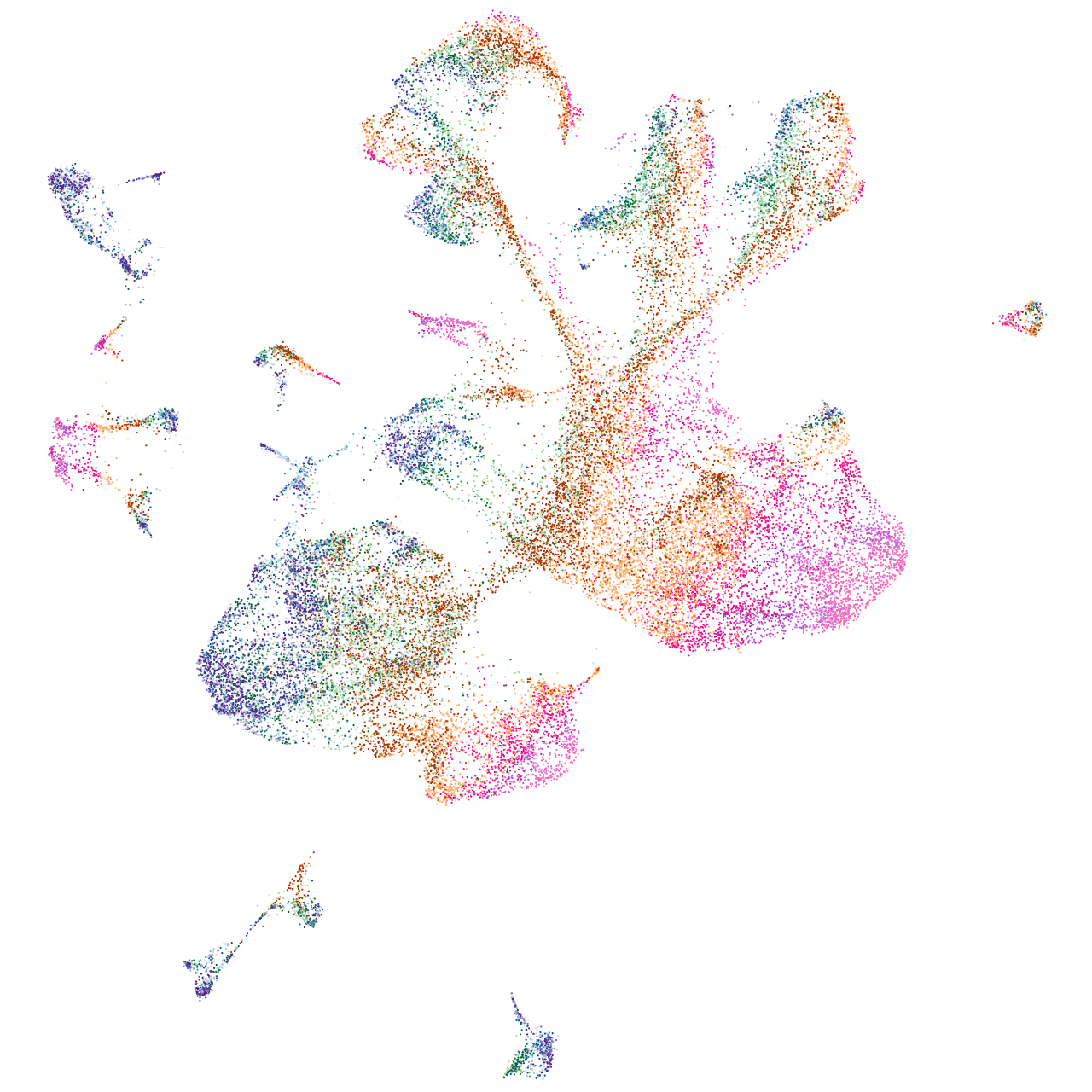
Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| pcna | 0.112 | ckbb | -0.077 |
| znfl2a | 0.111 | rtn1a | -0.063 |
| cdx4 | 0.107 | pvalb1 | -0.060 |
| fen1 | 0.105 | gpm6aa | -0.060 |
| rpa2 | 0.104 | pvalb2 | -0.059 |
| nasp | 0.103 | CU467822.1 | -0.057 |
| dut | 0.102 | hbbe1.3 | -0.055 |
| rrm1 | 0.101 | gpm6ab | -0.054 |
| selenoh | 0.101 | tmsb | -0.053 |
| chaf1a | 0.100 | rnasekb | -0.053 |
| npm1a | 0.100 | gapdhs | -0.052 |
| mcm7 | 0.100 | si:dkeyp-75h12.5 | -0.051 |
| CABZ01032351.1 | 0.098 | myt1b | -0.050 |
| slbp | 0.097 | actc1b | -0.050 |
| nutf2l | 0.096 | CR383676.1 | -0.050 |
| mcm5 | 0.096 | stmn1b | -0.049 |
| mcm6 | 0.095 | hbae3 | -0.049 |
| lig1 | 0.094 | elavl3 | -0.047 |
| anp32b | 0.093 | fabp7a | -0.047 |
| hspb1 | 0.093 | slc1a2b | -0.046 |
| apela | 0.093 | gng3 | -0.046 |
| rrm2 | 0.093 | CU634008.1 | -0.046 |
| mcm3 | 0.093 | marcksl1b | -0.045 |
| dkc1 | 0.092 | cotl1 | -0.045 |
| stmn1a | 0.092 | elavl4 | -0.045 |
| ranbp1 | 0.091 | mylpfa | -0.044 |
| cdca7a | 0.091 | ppdpfb | -0.044 |
| CABZ01075068.1 | 0.090 | calm1a | -0.043 |
| ssrp1a | 0.089 | fez1 | -0.043 |
| hmgb2a | 0.089 | tnrc6c1 | -0.043 |
| rfc4 | 0.089 | sncb | -0.043 |
| nop58 | 0.088 | COX3 | -0.042 |
| cdca7b | 0.088 | atp6v1e1b | -0.042 |
| mcm2 | 0.088 | vamp2 | -0.042 |
| cdc6 | 0.087 | hbae1.1 | -0.042 |




