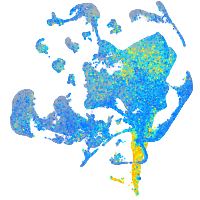
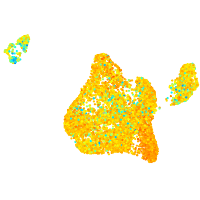
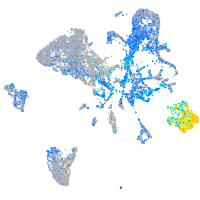
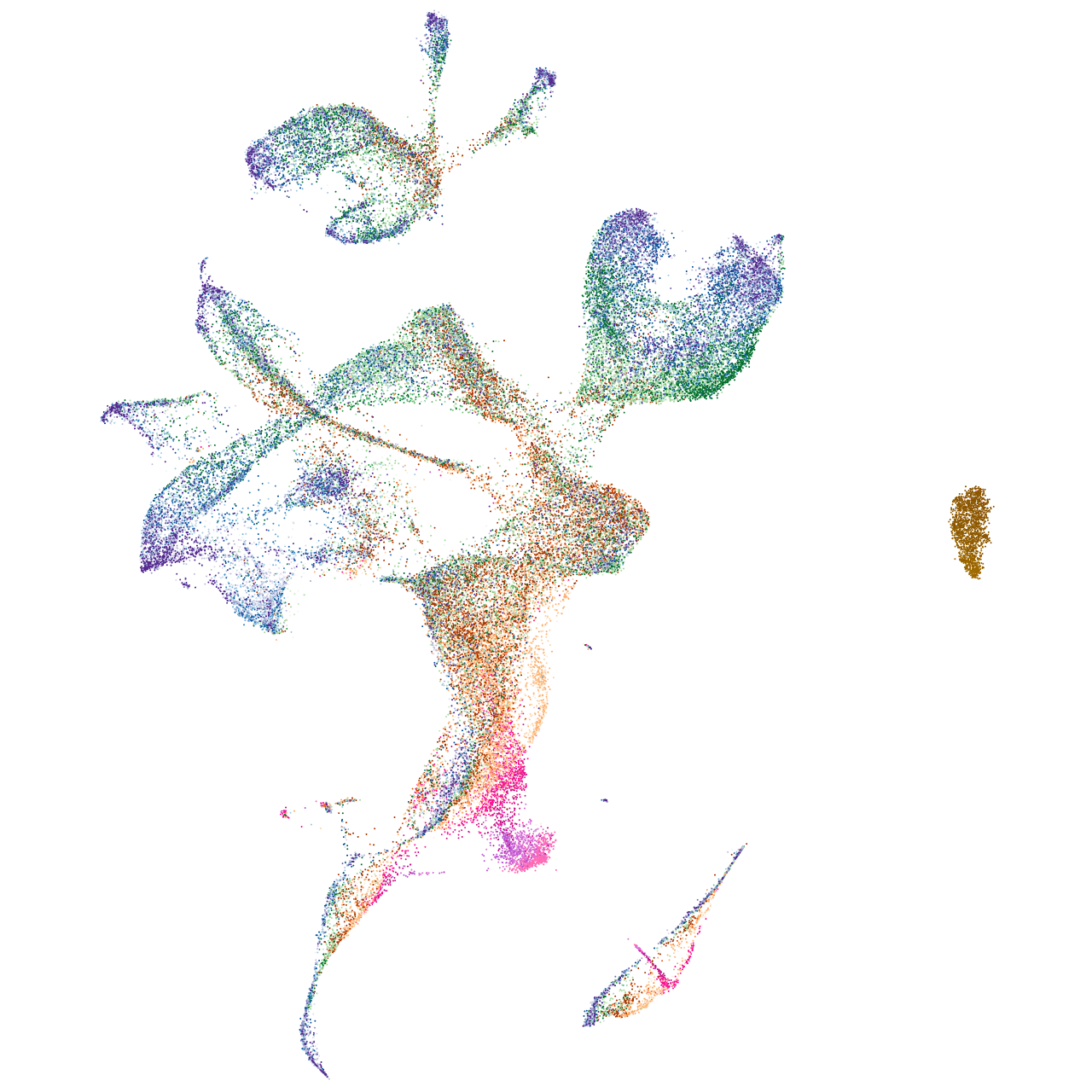heterogeneous nuclear ribonucleoprotein Ub
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| hmga1a | 0.393 | snap25b | -0.266 |
| si:ch211-222l21.1 | 0.389 | syt5b | -0.260 |
| seta | 0.357 | gapdhs | -0.260 |
| khdrbs1a | 0.352 | atp6v0cb | -0.248 |
| hnrnpabb | 0.351 | sypb | -0.240 |
| marcksb | 0.350 | atp6v1e1b | -0.228 |
| hmgb2b | 0.340 | ckbb | -0.226 |
| hnrnpa1b | 0.336 | cplx4a | -0.215 |
| hnrnpaba | 0.333 | atpv0e2 | -0.214 |
| h2afvb | 0.324 | rnasekb | -0.212 |
| cirbpa | 0.315 | sh3gl2a | -0.211 |
| ddx39ab | 0.312 | si:dkey-16p21.8 | -0.209 |
| setb | 0.307 | rs1a | -0.207 |
| cirbpb | 0.307 | aldocb | -0.205 |
| hnrnpa0b | 0.306 | tob1b | -0.203 |
| ran | 0.304 | tpi1b | -0.203 |
| snrpb | 0.303 | atp6v1g1 | -0.196 |
| nucks1a | 0.302 | atp6ap2 | -0.193 |
| syncrip | 0.301 | calb2b | -0.189 |
| cx43.4 | 0.300 | lin7a | -0.187 |
| hdac1 | 0.298 | gngt2b | -0.187 |
| top1l | 0.297 | pkma | -0.187 |
| rbm4.3 | 0.296 | eno1a | -0.186 |
| cbx3a | 0.295 | pvalb1 | -0.186 |
| rps6 | 0.293 | vamp1 | -0.182 |
| hmgn2 | 0.293 | CR383676.1 | -0.182 |
| si:ch211-288g17.3 | 0.292 | tob1a | -0.181 |
| ilf3b | 0.290 | nrn1lb | -0.179 |
| eef1a1l1 | 0.289 | gnb5b | -0.179 |
| eef2b | 0.289 | calm1b | -0.179 |
| shisa2a | 0.288 | pvalb2 | -0.178 |
| snrpd2 | 0.287 | CABZ01038709.1 | -0.175 |
| anp32b | 0.285 | mt-atp6 | -0.174 |
| ilf2 | 0.285 | rbp4l | -0.173 |
| snrpf | 0.284 | sncgb | -0.171 |