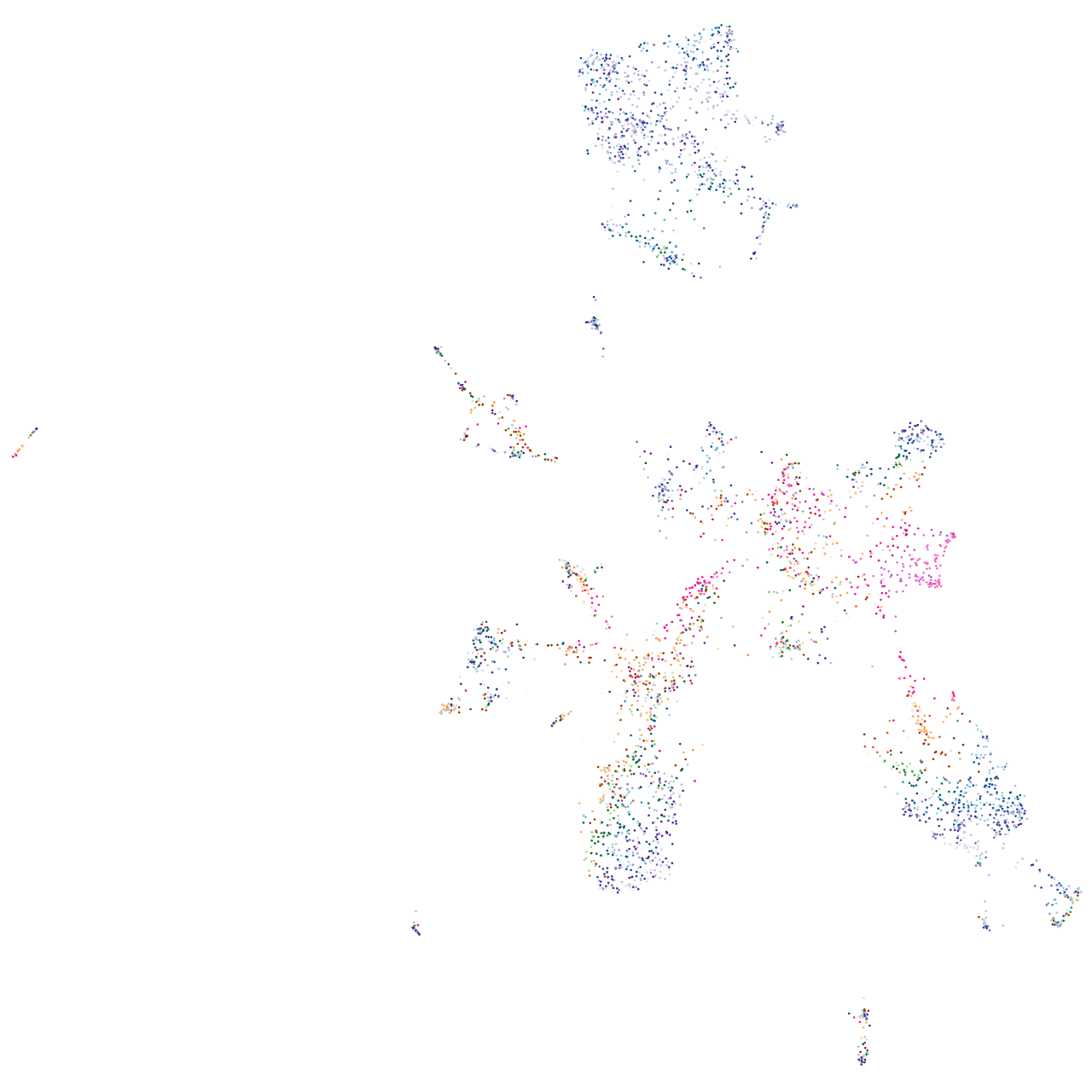
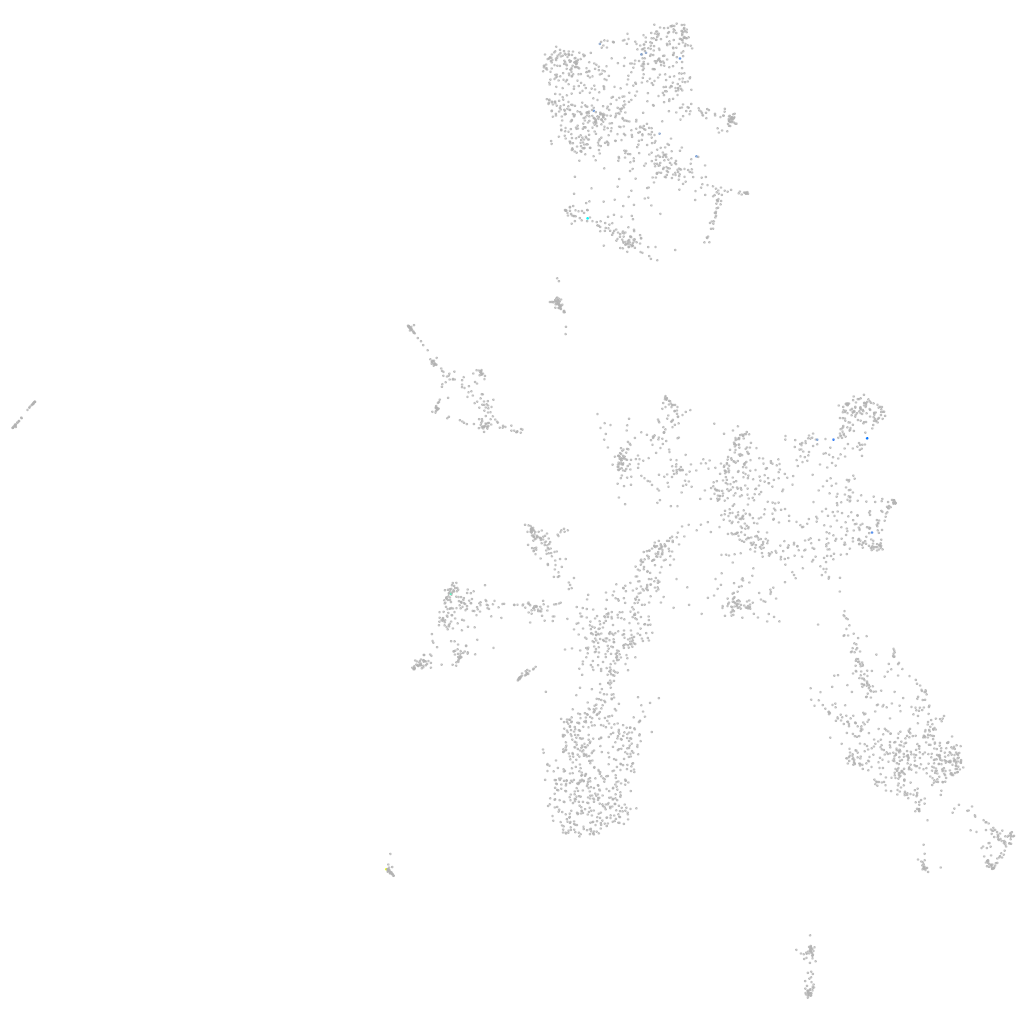
histidine ammonia-lyase
ZFIN 
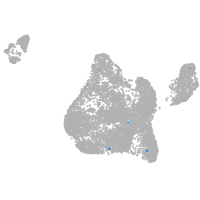
Other cell groups


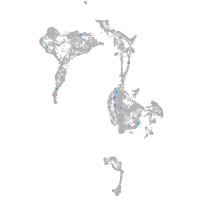
Expression by stage/cluster


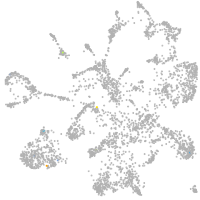
Correlated gene expression