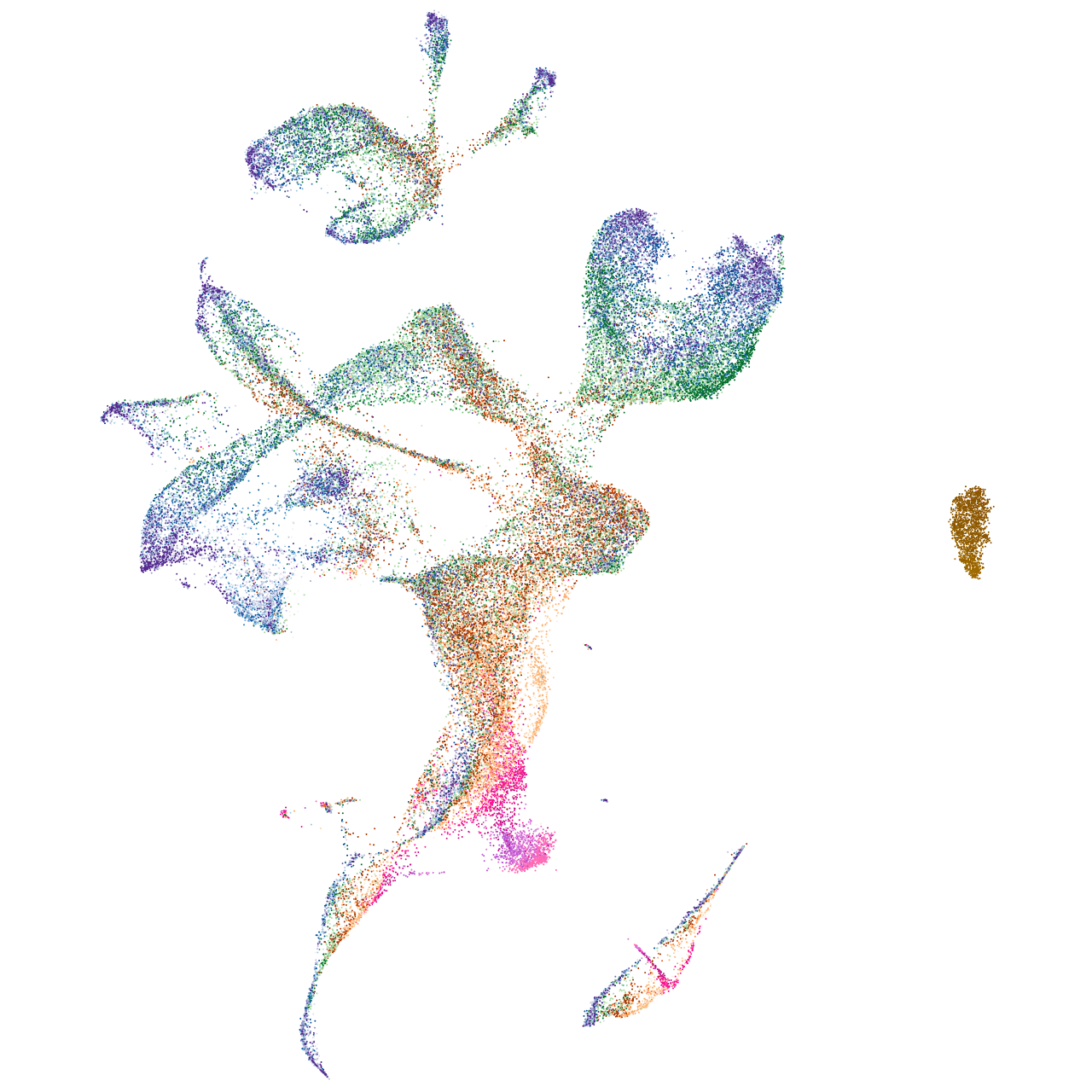FA complementation group B
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| mir125b-1 | 0.260 | ckbb | -0.045 |
| si:ch73-45o6.2 | 0.098 | atp6v0cb | -0.034 |
| dmc1 | 0.082 | atp6v1e1b | -0.032 |
| BX511253.1 | 0.082 | atp6v1g1 | -0.030 |
| si:dkey-28d5.4 | 0.078 | rnasekb | -0.029 |
| moto | 0.075 | snap25b | -0.029 |
| taar10 | 0.064 | ppdpfb | -0.028 |
| LOC103910488 | 0.064 | atpv0e2 | -0.028 |
| pcna | 0.063 | rtn1a | -0.028 |
| XLOC-007664 | 0.061 | calm1a | -0.028 |
| unga | 0.059 | gpm6aa | -0.027 |
| XLOC-039668 | 0.058 | sypb | -0.027 |
| stmn1a | 0.057 | vamp2 | -0.026 |
| FQ378040.1 | 0.057 | h2afx1 | -0.026 |
| lig1 | 0.056 | ndrg4 | -0.025 |
| CU207245.1 | 0.056 | h3f3c | -0.025 |
| zgc:110540 | 0.054 | scrt2 | -0.024 |
| fen1 | 0.054 | syt5b | -0.024 |
| ssrp1a | 0.053 | gng3 | -0.024 |
| esco2 | 0.053 | sncb | -0.024 |
| pole2 | 0.052 | gpm6ab | -0.024 |
| rpa2 | 0.052 | ppp1r14c | -0.023 |
| ftr99 | 0.052 | rs1a | -0.023 |
| dhfr | 0.051 | gapdhs | -0.023 |
| mcm7 | 0.051 | atp6ap2 | -0.023 |
| selenoh | 0.051 | pvalb2 | -0.023 |
| hirip3 | 0.051 | stmn1b | -0.023 |
| chaf1a | 0.051 | pvalb1 | -0.022 |
| dut | 0.051 | atp1a3a | -0.022 |
| zgc:113984 | 0.050 | oaz1b | -0.022 |
| ccnd1 | 0.050 | gng13b | -0.022 |
| banf1 | 0.050 | CR383676.1 | -0.022 |
| rfc5 | 0.050 | pcbp3 | -0.021 |
| rpa3 | 0.050 | eif1b | -0.021 |
| CU041363.1 | 0.049 | elavl3 | -0.021 |