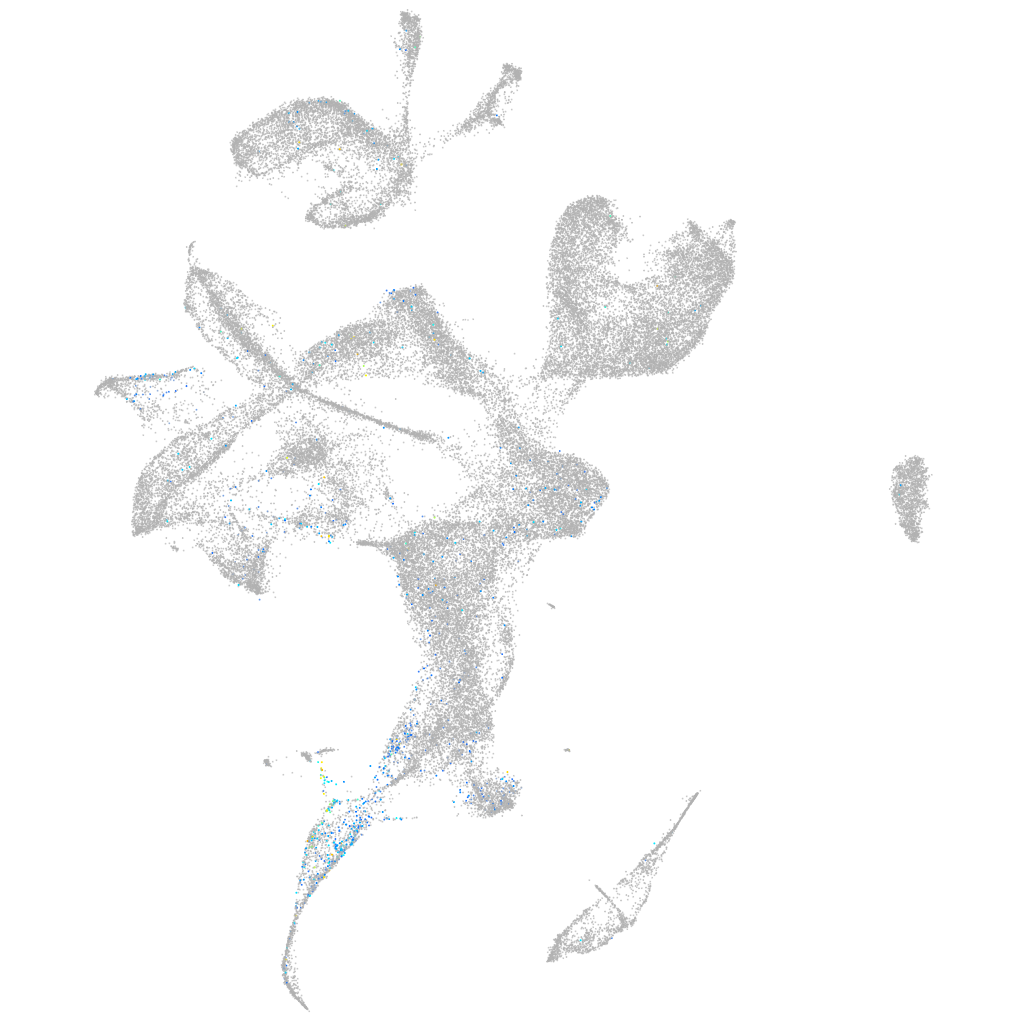
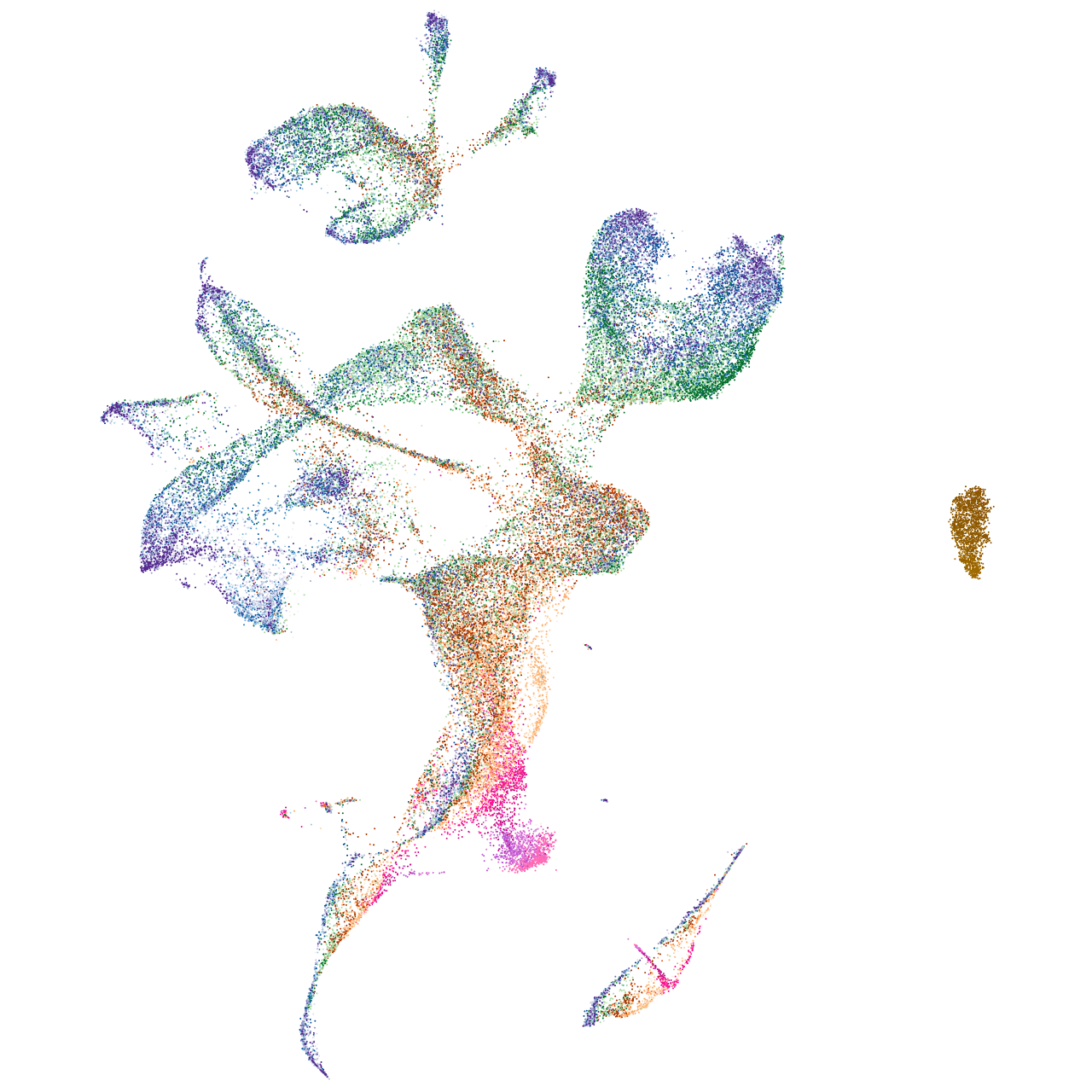enolase 4
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| smoc2 | 0.220 | ptmab | -0.061 |
| cyp27c1 | 0.198 | si:ch73-1a9.3 | -0.058 |
| vax1 | 0.192 | gpm6aa | -0.053 |
| tfec | 0.189 | gpm6ab | -0.048 |
| slc45a2 | 0.177 | ckbb | -0.048 |
| rgmd | 0.177 | ndrg4 | -0.044 |
| tmem88b | 0.171 | snap25b | -0.043 |
| dct | 0.171 | calb2b | -0.042 |
| ntn1a | 0.170 | vsx1 | -0.042 |
| tspan36 | 0.164 | foxg1b | -0.041 |
| pmela | 0.162 | anp32e | -0.041 |
| col9a1b | 0.162 | cspg5a | -0.040 |
| c1qtnf12 | 0.158 | pcbp3 | -0.038 |
| sytl2b | 0.156 | epb41a | -0.037 |
| tyrp1b | 0.154 | tuba1c | -0.037 |
| sfrp1a | 0.152 | marcksl1b | -0.037 |
| gstp1 | 0.148 | syt5b | -0.037 |
| gpr143 | 0.147 | mpp6b | -0.037 |
| mitfa | 0.147 | hmgb1a | -0.037 |
| fabp11a | 0.147 | hmgn6 | -0.036 |
| rab38 | 0.144 | atp1b2a | -0.036 |
| agtrap | 0.144 | gapdhs | -0.035 |
| tyrp1a | 0.144 | nova2 | -0.035 |
| FP085398.1 | 0.143 | cadm3 | -0.035 |
| foxi2 | 0.142 | scrt2 | -0.035 |
| cldn19 | 0.142 | gng13b | -0.034 |
| tspan10 | 0.141 | rnasekb | -0.034 |
| trpm1b | 0.141 | neurod4 | -0.034 |
| qdpra | 0.140 | pcp4l1 | -0.033 |
| pcbd1 | 0.138 | bambia | -0.033 |
| c1qtnf1 | 0.137 | atp1a3a | -0.033 |
| zgc:110591 | 0.135 | efnb2a | -0.033 |
| ldlrap1a | 0.134 | olfm1b | -0.033 |
| foxp4 | 0.133 | gnb3a | -0.033 |
| fgfrl1a | 0.132 | atp6v0cb | -0.033 |