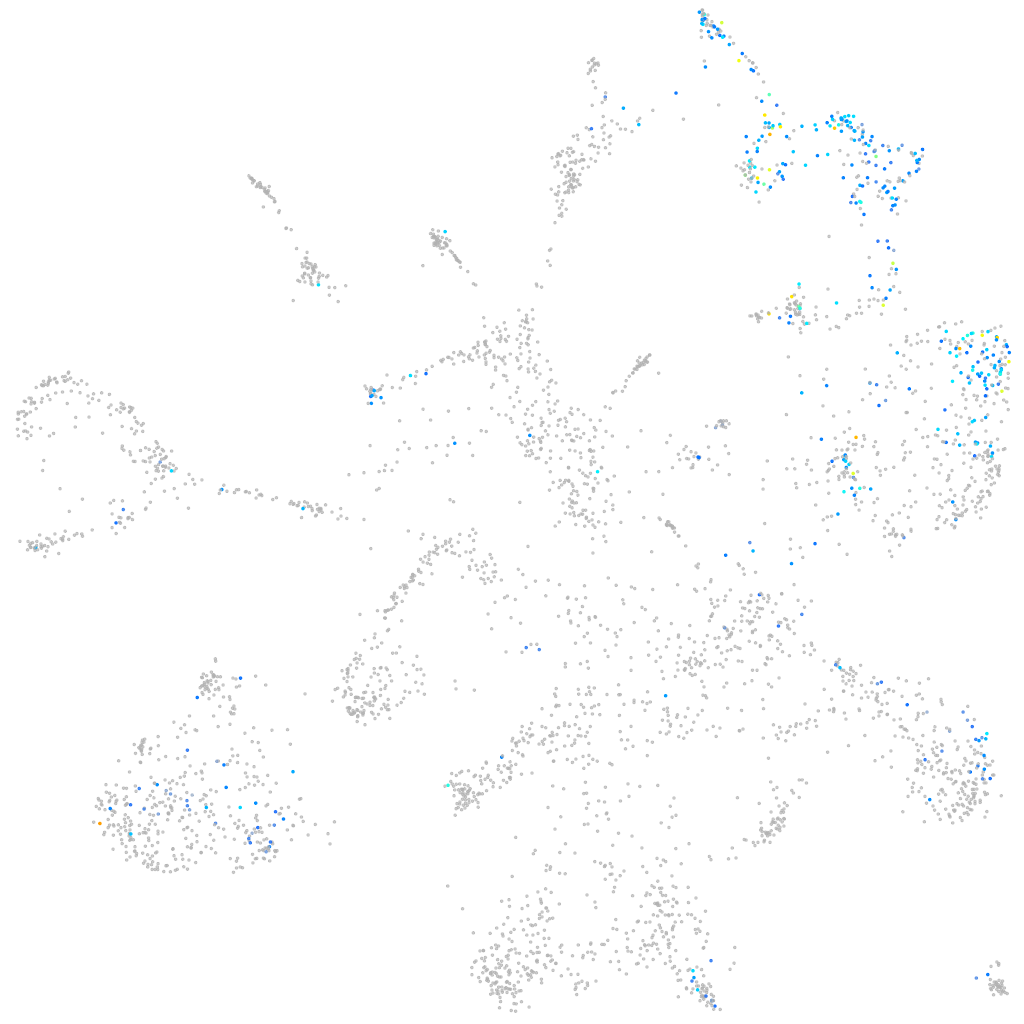
CXADR Ig-like cell adhesion molecule
ZFIN 
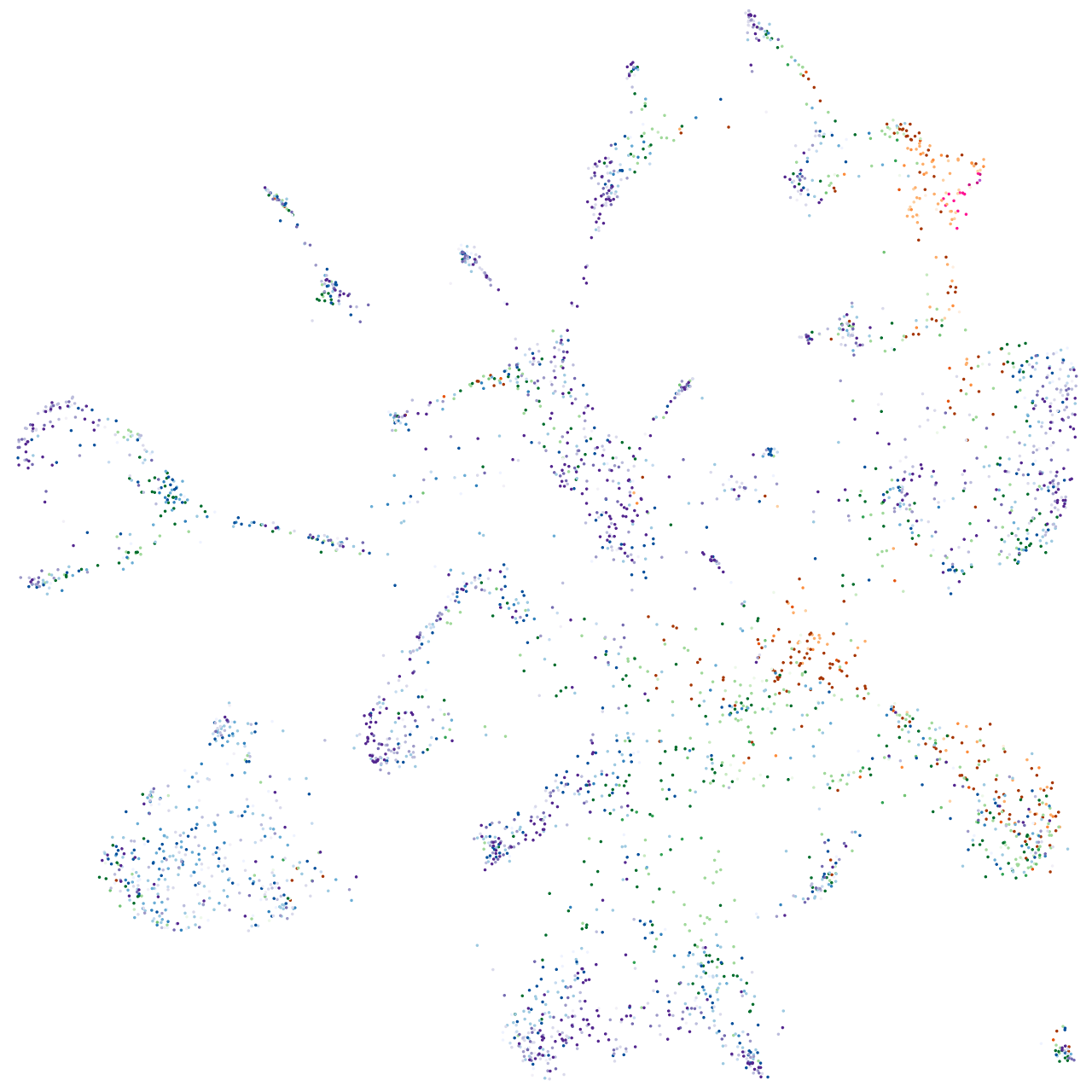
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| mmel1 | 0.396 | BX088707.3 | -0.225 |
| podxl | 0.393 | tpm1 | -0.195 |
| gata5 | 0.345 | acta2 | -0.181 |
| krt94 | 0.342 | myl9a | -0.179 |
| tmem88b | 0.342 | tagln | -0.168 |
| jam2b | 0.341 | myl6 | -0.151 |
| aqp1a.1 | 0.329 | myh11a | -0.146 |
| bnc2 | 0.327 | ptmaa | -0.137 |
| gstm.3 | 0.323 | lmod1b | -0.137 |
| nr0b2a | 0.318 | nkx2.3 | -0.137 |
| fabp11a | 0.315 | mylkb | -0.137 |
| slc29a1b | 0.304 | inka1a | -0.135 |
| bcam | 0.302 | foxf1 | -0.134 |
| COLEC10 | 0.299 | fhl2a | -0.126 |
| crb2a | 0.298 | gapdhs | -0.124 |
| krt8 | 0.294 | zgc:153704 | -0.124 |
| sfrp5 | 0.291 | XLOC-025423 | -0.124 |
| rhag | 0.286 | rbpms2a | -0.122 |
| synpo2lb | 0.283 | tpm2 | -0.121 |
| hspb1 | 0.282 | ckbb | -0.120 |
| arl4aa | 0.274 | si:ch211-62a1.3 | -0.120 |
| cd151 | 0.273 | ppiab | -0.119 |
| lbx2 | 0.270 | cap1 | -0.118 |
| alpi.1 | 0.270 | calm2a | -0.113 |
| gata6 | 0.268 | XLOC-041870 | -0.112 |
| colec12 | 0.261 | gpia | -0.111 |
| wt1b | 0.259 | ntn5 | -0.111 |
| lmod2a | 0.255 | igfbp7 | -0.111 |
| tbx1 | 0.252 | csrp1b | -0.109 |
| tgm2b | 0.251 | mcamb | -0.108 |
| angptl6 | 0.251 | mdkb | -0.108 |
| lrp2a | 0.244 | desmb | -0.107 |
| ftr82 | 0.239 | cald1b | -0.106 |
| cav1 | 0.238 | pdgfrb | -0.106 |
| krt18a.1 | 0.235 | myocd | -0.104 |