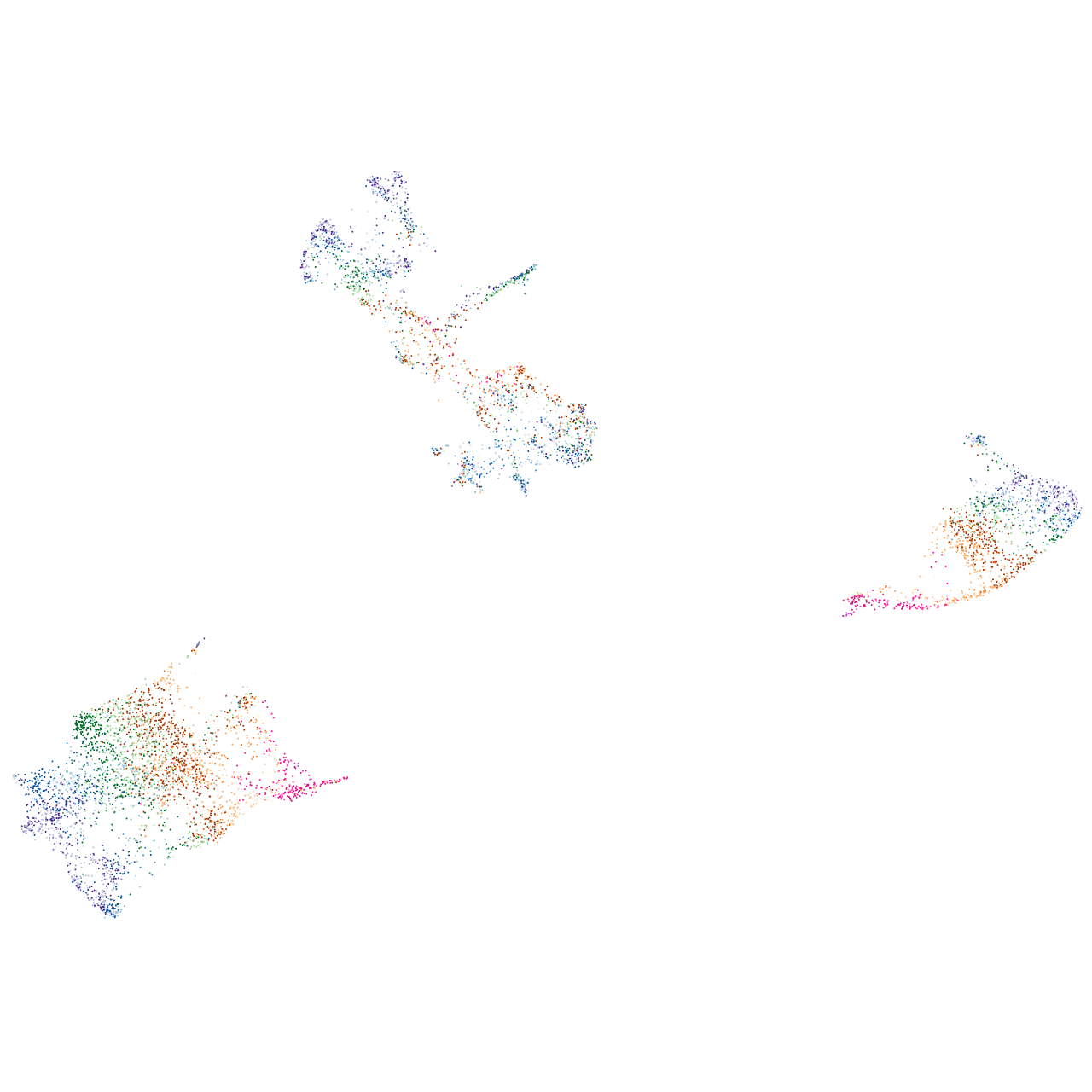
connector enhancer of kinase suppressor of Ras 2b
ZFIN 
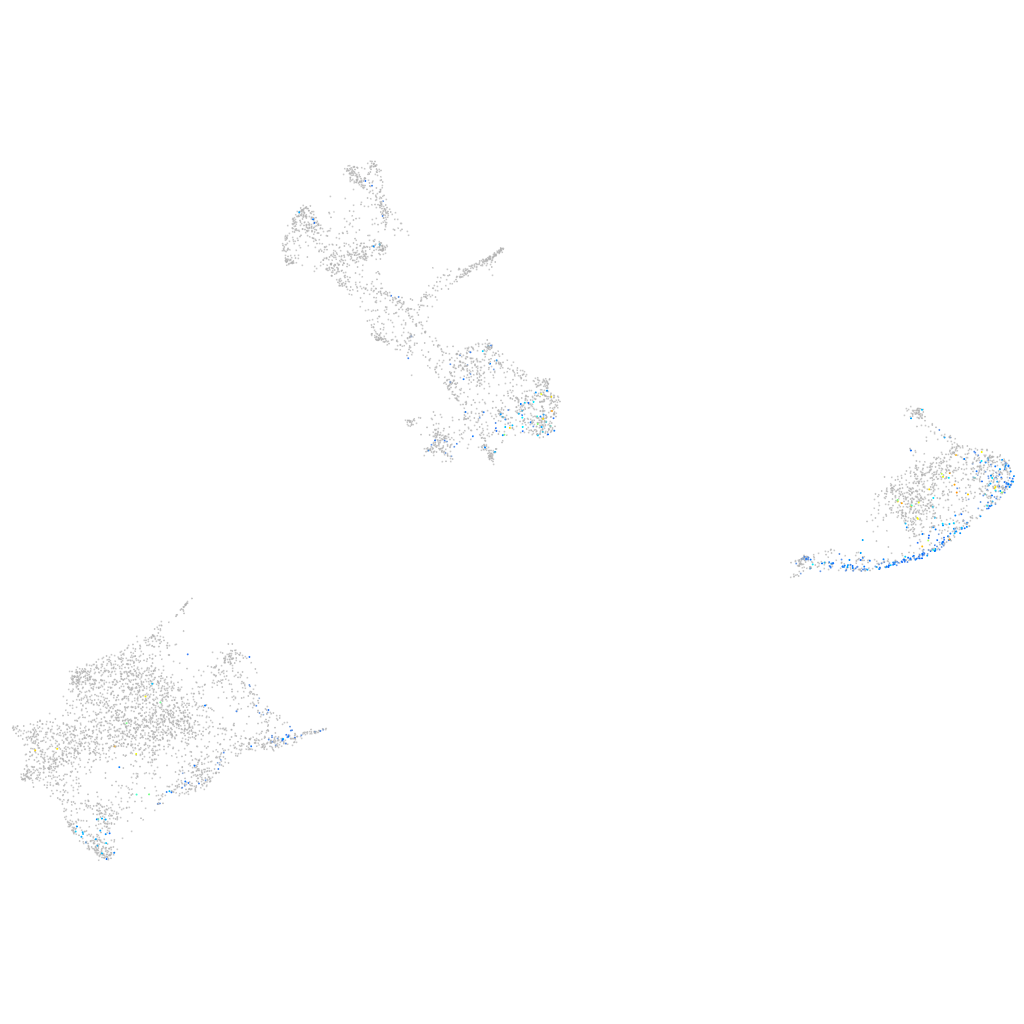
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| lamp1a | 0.252 | paics | -0.142 |
| kita | 0.243 | uraha | -0.131 |
| SPAG9 | 0.231 | CABZ01021592.1 | -0.121 |
| oca2 | 0.229 | mdh1aa | -0.118 |
| slc24a5 | 0.228 | si:dkey-251i10.2 | -0.106 |
| tyrp1a | 0.226 | atic | -0.098 |
| dct | 0.222 | tmem130 | -0.088 |
| tyr | 0.222 | impdh1b | -0.088 |
| pmela | 0.219 | defbl1 | -0.087 |
| tyrp1b | 0.215 | slc2a15a | -0.086 |
| slc24a4a | 0.201 | apoda.1 | -0.085 |
| zgc:91968 | 0.200 | phyhd1 | -0.084 |
| ctsc | 0.194 | prdx1 | -0.080 |
| tmem243b | 0.194 | gpnmb | -0.080 |
| aadac | 0.194 | prps1a | -0.080 |
| mitfa | 0.192 | si:ch211-256m1.8 | -0.080 |
| kcnj13 | 0.190 | si:dkey-197i20.6 | -0.077 |
| si:ch73-389b16.1 | 0.189 | cyb5a | -0.076 |
| mtbl | 0.187 | prdx5 | -0.075 |
| gpr61 | 0.187 | krt18b | -0.074 |
| tfap2e | 0.184 | lypc | -0.073 |
| myo9ab | 0.183 | si:ch211-251b21.1 | -0.073 |
| prkar1b | 0.181 | hmgb2a | -0.073 |
| si:ch211-195b13.1 | 0.181 | alx4a | -0.072 |
| atp6v0a2b | 0.180 | bco1 | -0.072 |
| rabl6b | 0.178 | unm-sa821 | -0.071 |
| mchr2 | 0.178 | rbp4l | -0.071 |
| slc7a8a | 0.177 | akap12a | -0.068 |
| slc45a2 | 0.176 | pltp | -0.067 |
| slc7a5 | 0.176 | aqp1a.1 | -0.067 |
| slc29a3 | 0.175 | pvalb2 | -0.067 |
| slc3a2a | 0.175 | sult1st1 | -0.066 |
| slc37a2 | 0.174 | si:ch73-89b15.3 | -0.066 |
| slc39a10 | 0.173 | actc1b | -0.065 |
| msx1b | 0.173 | alx4b | -0.064 |