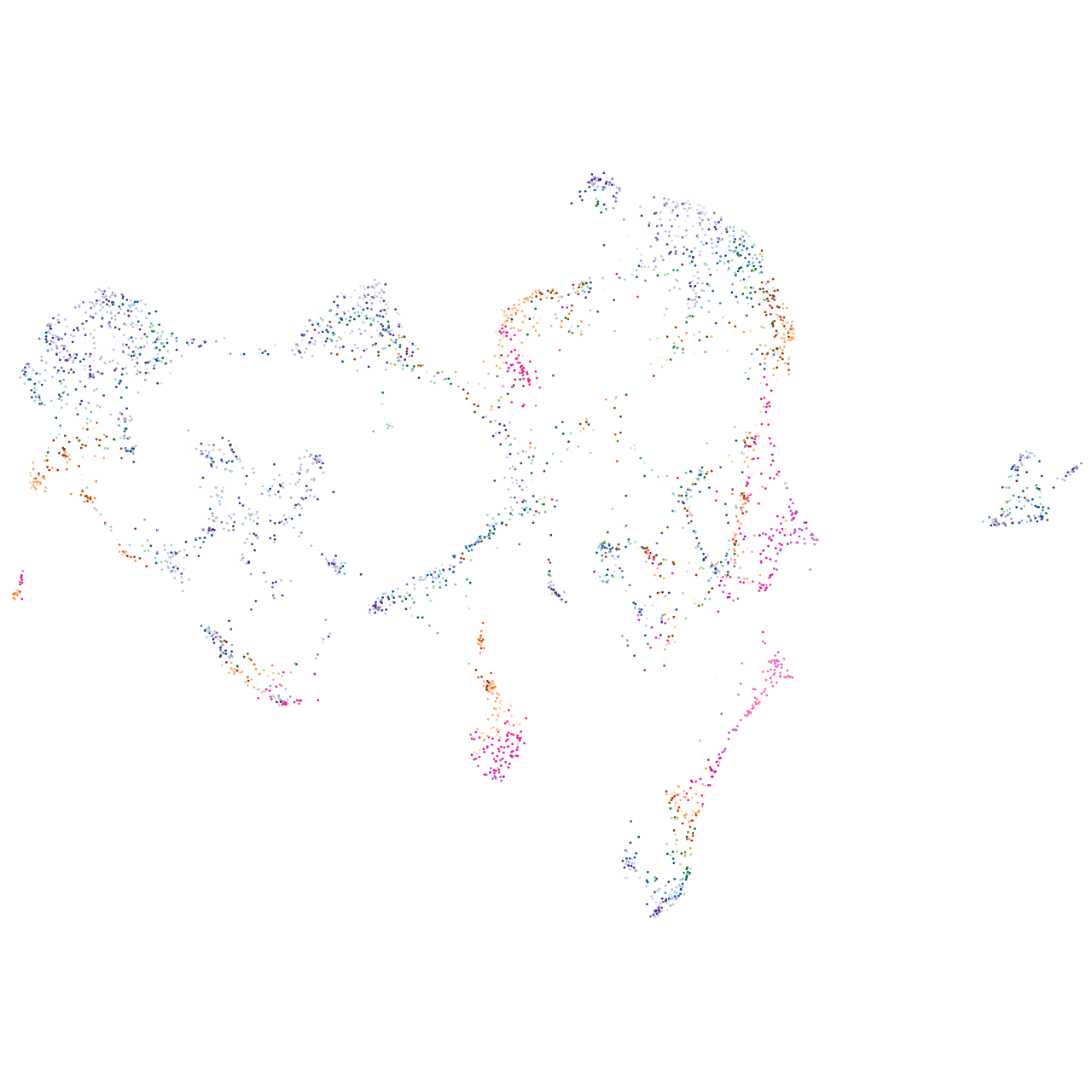
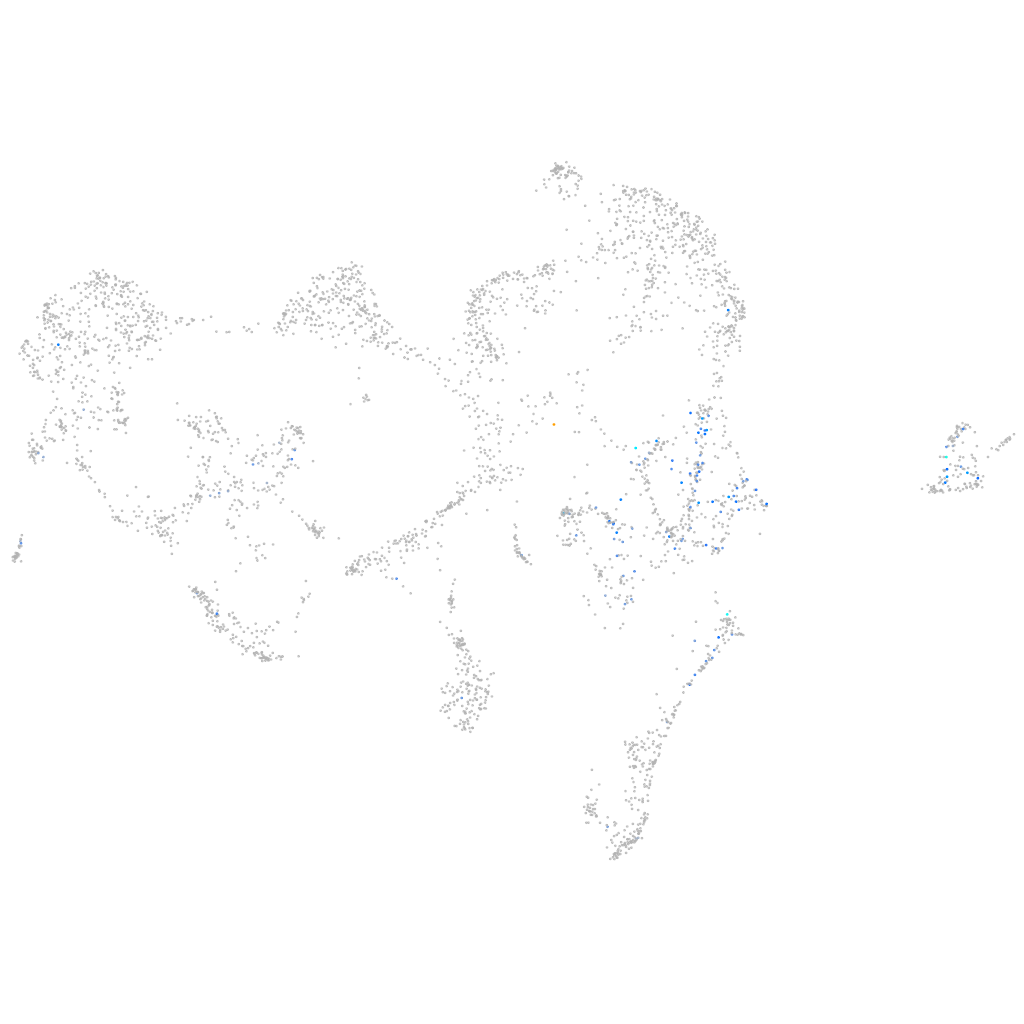
cell division cycle 7 homolog (S. cerevisiae)
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| nasp | 0.257 | si:ch211-195b11.3 | -0.122 |
| dut | 0.251 | selenow1 | -0.119 |
| dhfr | 0.245 | BX000438.2 | -0.112 |
| chaf1a | 0.245 | si:dkey-193p11.2 | -0.111 |
| lig1 | 0.244 | icn2 | -0.104 |
| rbbp4 | 0.241 | soul2 | -0.103 |
| zgc:110540 | 0.239 | tob1b | -0.100 |
| cacna2d2a | 0.237 | selenow2b | -0.099 |
| rrm1 | 0.236 | ppdpfb | -0.098 |
| slbp | 0.235 | icn | -0.098 |
| LOC110439438 | 0.230 | si:dkey-248g15.3 | -0.098 |
| pcna | 0.225 | zgc:110333 | -0.097 |
| rpa2 | 0.221 | zgc:153284 | -0.096 |
| stmn1a | 0.217 | abcb5 | -0.095 |
| gins3 | 0.216 | aldob | -0.093 |
| chaf1b | 0.216 | zgc:193505 | -0.092 |
| hmgb1b | 0.216 | enosf1 | -0.091 |
| rrm2 | 0.215 | ppl | -0.090 |
| mcm7 | 0.213 | gstp1 | -0.090 |
| lpcat2 | 0.209 | rplp2 | -0.089 |
| dek | 0.208 | si:dkey-247k7.2 | -0.089 |
| hmgb2b | 0.203 | COX3 | -0.088 |
| six6a | 0.203 | ahnak | -0.088 |
| ccna2 | 0.203 | mgst1.2 | -0.087 |
| CABZ01005379.1 | 0.200 | si:ch73-359m17.9 | -0.085 |
| ccne2 | 0.198 | nfe2l2a | -0.085 |
| rfc3 | 0.198 | krt17 | -0.085 |
| vrk1 | 0.198 | rnf128a | -0.084 |
| fam161a | 0.197 | wu:fb18f06 | -0.083 |
| mcm3 | 0.196 | scel | -0.082 |
| cdca5 | 0.194 | prdx5 | -0.082 |
| haus4 | 0.194 | tmem59 | -0.082 |
| supt16h | 0.193 | krt91 | -0.081 |
| TIMM22 | 0.193 | spint2 | -0.079 |
| prim2 | 0.193 | atp1b1b | -0.079 |