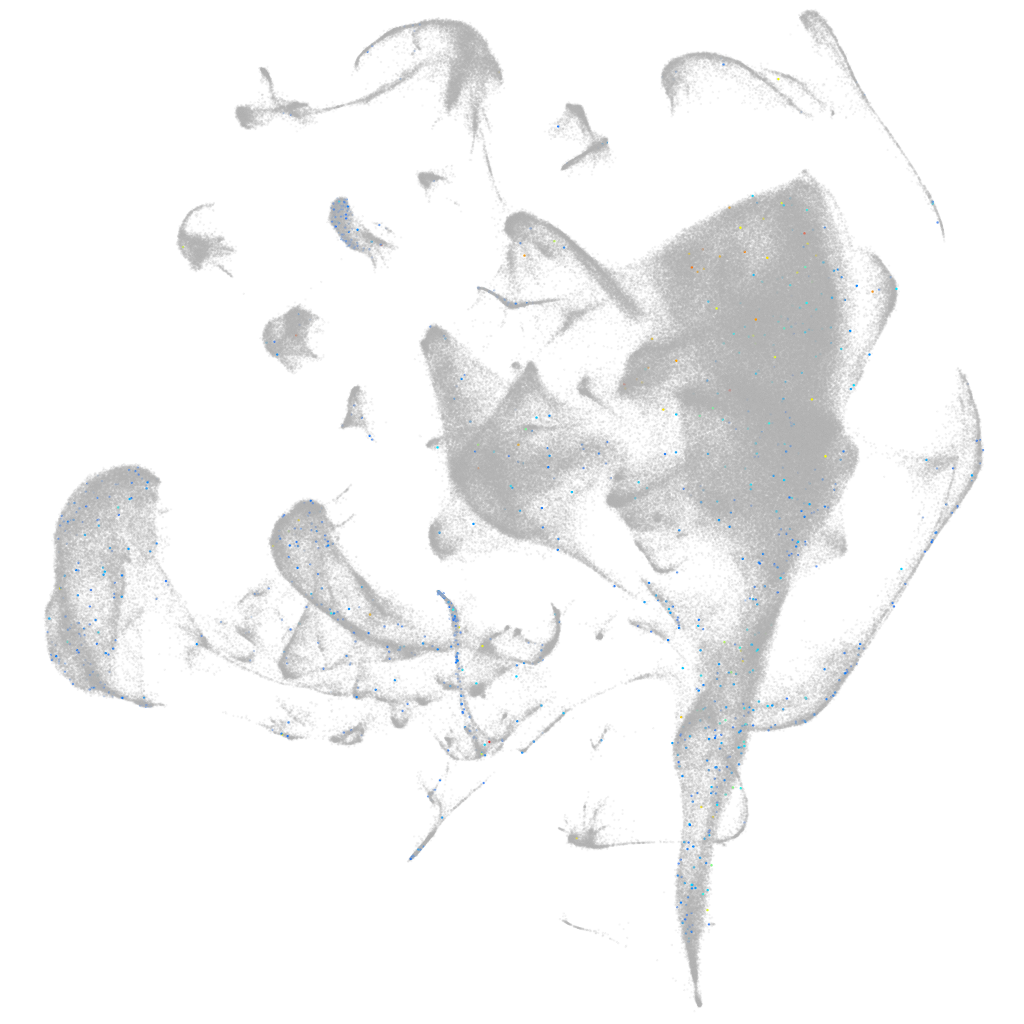
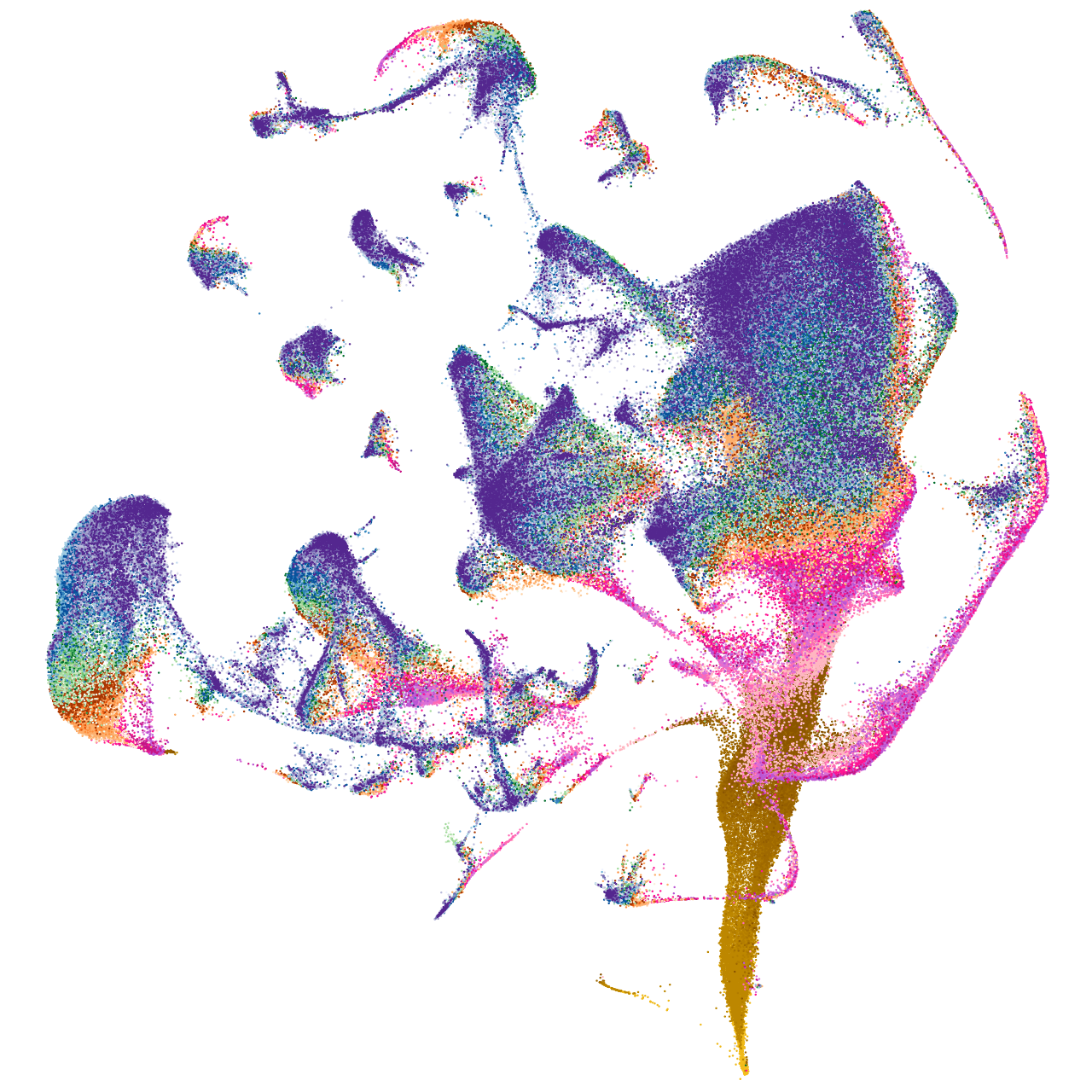Other cell groups


all cells |
blastomeres |
PGCs |
endoderm |
axial mesoderm |
muscle  |
mural cells / non-skeletal muscle  |
mesenchyme  |
hematopoietic / vasculature  |
pronephros  |
neural |
spinal cord / glia |
eye |
taste / olfactory |
otic / lateral line |
epidermis |
periderm |
fin |
ionocytes / mucous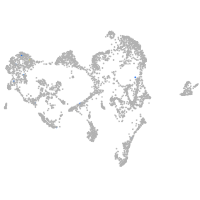 |
pigment cells |
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| cyp8b1 | 0.086 | marcksl1a | -0.027 |
| si:ch211-117n7.7 | 0.086 | tuba1c | -0.026 |
| rdh1 | 0.085 | gpm6aa | -0.024 |
| creb3l3a | 0.084 | nova2 | -0.024 |
| pla2g12b | 0.084 | mdkb | -0.021 |
| golt1a | 0.083 | atp6v0cb | -0.020 |
| rltgr | 0.083 | gpm6ab | -0.020 |
| sult3st3 | 0.083 | rtn1a | -0.020 |
| ace2 | 0.082 | stmn1b | -0.020 |
| aqp8a.2 | 0.082 | tuba1a | -0.020 |
| si:dkey-30c15.13 | 0.082 | ckbb | -0.019 |
| sult3st2 | 0.082 | hbbe1.3 | -0.019 |
| tmem86b | 0.082 | hmgb1b | -0.019 |
| CU682777.2 | 0.081 | rnasekb | -0.019 |
| CR855996.2 | 0.080 | celf2 | -0.018 |
| dnajc22 | 0.080 | elavl3 | -0.018 |
| wu:fa56d06 | 0.080 | gapdhs | -0.018 |
| zanl | 0.080 | hbae3 | -0.018 |
| tm4sf5 | 0.080 | jpt1b | -0.018 |
| cpo | 0.079 | actc1b | -0.017 |
| slc6a19a.2 | 0.079 | cspg5a | -0.017 |
| ugt5b3 | 0.079 | epb41a | -0.017 |
| zgc:77938 | 0.079 | fam168a | -0.017 |
| cyp7a1 | 0.078 | hmgb3a | -0.017 |
| fbp1b | 0.078 | pvalb1 | -0.017 |
| mbl2 | 0.078 | pvalb2 | -0.017 |
| mttp | 0.078 | atpv0e2 | -0.016 |
| AL831745.1 | 0.077 | cadm3 | -0.016 |
| cyp2x9 | 0.077 | fabp7a | -0.016 |
| si:ch211-77g15.32 | 0.077 | gng3 | -0.016 |
| si:dkeyp-73b11.8 | 0.077 | hbae1.1 | -0.016 |
| ugt1a7 | 0.077 | sncb | -0.016 |
| zgc:110410 | 0.077 | elavl4 | -0.015 |
| zgc:153968 | 0.077 | fabp3 | -0.015 |
| cx28.9 | 0.077 | gnao1a | -0.015 |